Whilst most molecular HLA typing methodologies remain relatively static, next-generation sequencing (NGS) has tended to evolve quickly as product manufacturers respond to user suggestions, reagent development and changes in sequencing technology. One Lambda released AllType™ in Summer 2017: a NGS workflow for all loci, all platforms, all in a single reaction. This NGS workflow was a game changer, beginning with the long-range PCR of all 11 classical HLA genes in a single multiplexed reaction. Whilst AllType™ remains a competitive workflow, One Lambda have now released an alternative NGS solution: AllType™ FASTplex™ NGS.
AllType™ FASTplex™ NGS
The rationale behind AllType™ FASTplex™ NGS was to simplify the NGS workflow, save time, and yet still generate high-quality results. FASTplex™ sets a new standard of simplicity for NGS and is a simple assay that anyone can learn. Like the classic AllType™ workflow, FASTplex™ begins with all 11 classical HLA loci (HLA-A,B,C,DRB1,DRB3/4/5,DQA1,DQB1,DPA1,DPB1) co-amplified in a single multiplexed reaction (Table 1). After an initial thermal cycling time of only 2.5 hours, patient-specific amplicons are pooled into a single tube within an hour of commencing library preparation. This means that approximately 70% of the FASTplex™ protocol is single tube workflow. The consequences of this are significant: from DNA to sequencing, sample preparation takes less than seven hours and requires only 90 minutes of hands-on time. An easy single-tube workflow reduces the chances of pipetting error and means that automation isn’t required. By eliminating most of the pipetting, purification and quantification steps required by ‘traditional’ methods, FASTplex™ significantly reduced the consumables required. Compared to the classic AllType™ workflow, FASTplex™ uses less than half the plastics, pipette tips and consumables: good news for both operational costs and the environment. The AllType™ FASTplex™ NGS product also includes paramagnetic beads, further decreasing the cost per test resulting from consumables and third-party reagents.
Table 1: Gene content and coverage for AllType™ FASTplex™ NGS
| Loci | Coverage |
|---|---|
| HLA-A | Whole gene |
| HLA-B | Whole gene |
| HLA-C | Whole gene |
| HLA-DRB1 | Exon 1 + Exon 2 – Intron 5 |
| HLA-DRB345 | Exon 2 – Intron 5 |
| HLA-DQB1 | Exon 1 + Exon 2 – 3’ UTR |
| HLA-DQA1 | Whole gene |
| HLA-DPB1 | Exon 1 + Exon 2 – 3’ UTR |
| HLA-DPA1 | Whole gene |
The FASTplex™ Workflow
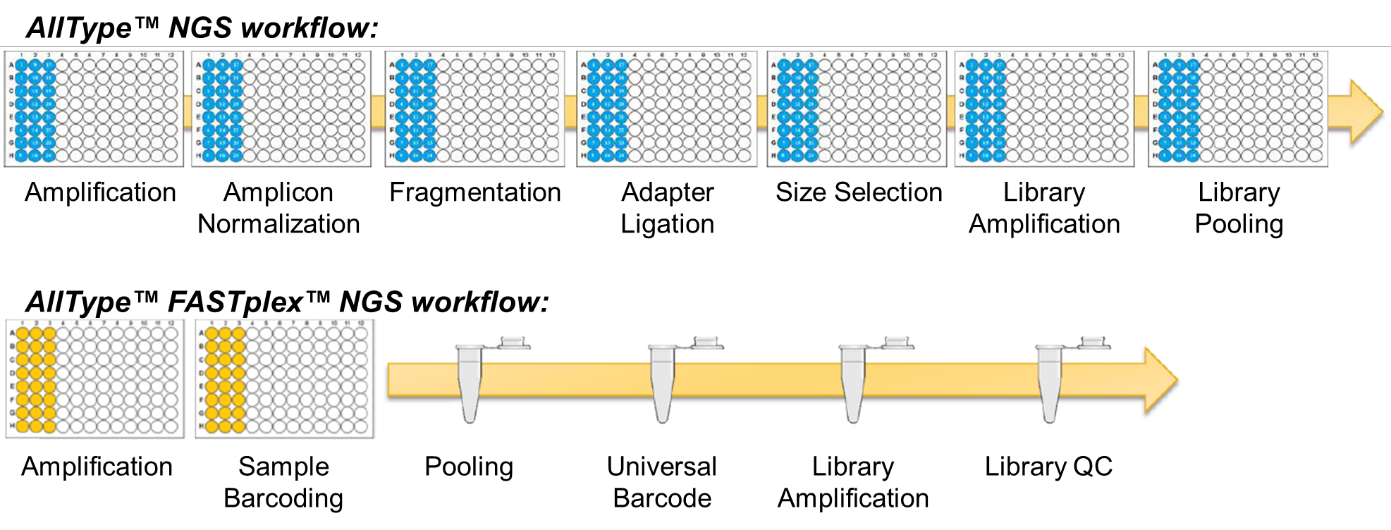
The AllType™ and AllType™ FASTplex™ workflows are compared in Figure 1. FASTplex™ builds on the classic workflow and removes some elements that might be considered undesirable in a non-automated laboratory. Following amplification and paramagenetic bead purification, sample-specific amplicons are quantified using a Qubit instrument. AllType™ requires sample-specific normalisation to proceed with 100 ng DNA whilst FASTplex™ allows the user to proceed with 3-30 ng DNA, reducing the pipetting burden as samples ought to fall within this range. A sophisticated workbook-based tool guides the user through the FASTplex™ workflow, indicating if adjustments are required and performing the necessary calculations. Unlike the classic workflow where fragmentation, adapter ligation (‘barcoding’) and size-selection are discrete processes, the FASTplex™ workflow simultaneously fragments and barcodes amplicons using a set of ‘FAST’ reagents pre-plated in a 96-well tray; size-selection is not required. At this point FASTplex™ samples are pooled and the remainder of the workflow occurs in a single tube. In the FASTplex™ workflow an additional ‘universal barcode’ is ligated to all fragments prior to a secondary PCR amplification. Thereafter the library is cleaned using paramagnetic beads, quantified and diluted to the concentration required for sequencing. Working in a single-tube significantly increases the speed and simplicity of the workflow; although bead-based purification and quantitation is still required the volume is significantly reduced compared to e.g. 24 samples.
Figure 1: The AllType™ and AllType™ FASTplex™ NGS workflows compared.
Multi-Platform Support
Both AllType™ and AllType™ FASTplex™ are supported on multiple Illumina platforms and kits, in addition to Ion Torrent platforms. This provides verified flexibility for the end user, who may choose to utilise a sequencing platform shared by multiple users or acquire their own ‘entry level’ sequencer (e.g. the iSeq 100, with a purchase price of approximately £20,000). Many sequencing platforms also offer different ‘kit’ sizes which allow users to implement a solution appropriate to their laboratory, i.e. balancing throughput, ‘batch’ cost per sample (considering the sequencing kit) and turnaround time.
Table 2: Illumina sequencer throughput for AllType™ and AllType™ FASTplex™.
| Platform | Kit | Throughput | Sequencing Time |
|---|---|---|---|
| MiSeq | Standard | ≤ 96 samples | 24 hrs |
| Micro | ≤ 24 samples | 19 hrs | |
| Nano | ≤ 8 samples | 17 hrs | |
| MiniSeq | High-Output | ≤ 96 samples | 24 hrs |
| Mid-Output | ≤ 72 samples | 17 hrs | |
| iSeq 100 | Reagent | ≤ 24 samples | 17.5 hrs |
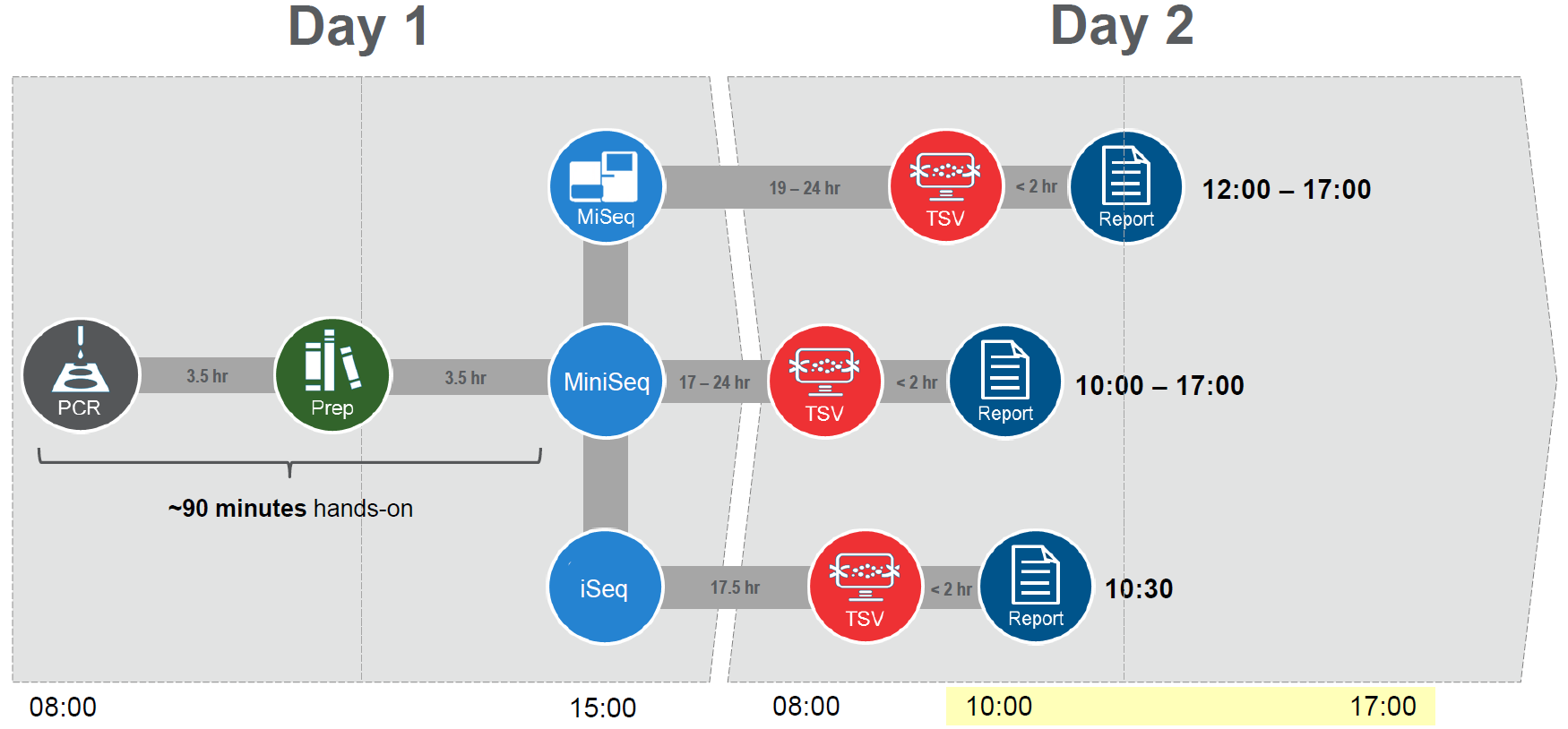
The sequencing platform and kit used also impacts the sequencing time and thus the time taken from sample to HLA genotyping results. Using the fastest solution, AllType™ FASTplex™ allows the user to obtain HLA genotyping results a mere 26 hours after starting their testing (Figure 2).
Figure 2: AllType™ FASTplex™ is the fastest sample-to-result NGS workflow for HLA.
TypeStream Visual™ 3.1: Fast, Fully-Automated Analysis
AllType™ FASTplex™ is supported by TypeStream Visual™ 3.1: intelligent software for the analysis of AllType™ data from Ion Torrent, Illumina and PacBio sequencing platforms. TypeStream Visual ™ 3.1 retains the familiar (Fusion-esque) ‘front-end’ environment but builds on LIMS integration: SystemLink offer a HistoTrac module that interfaces directly with the TypeStream Visual™ database, and HLA typing data can now be exported to One Lambda’s HLA Fusion™ software with only three clicks. This allows the integration of high-resolution genotypes with patient/donor records and facilitates antibody tracking and epitope-level analysis using HLAMatchmaker functionality built into HLA Fusion™.
The TypeStream Visual™ auto-analysis feature streamlines data migration so that data are analysed even when the user is away from the laboratory. TypeStream Visual™ 3.1 also features an analysis engine that allows multiple samples to be analysed simultaneously, resulting in faster analysis times. This ensures that AllType™ and AllType™ FASTplex™ are the fastest sample-to-result NGS workflows for HLA genotyping. TypeStream Visual™ 3.1 offers more user-definable features including health metrics, system comments and the ability to exclude specific gene regions (e.g. micro-satellites). The analysis experience has been redesigned with a streamlined interface: fewer ‘levels’ of analysis require fewer ‘clicks’ to assign a result (Figure 3). Upgrading to and maintaining TypeStream Visual™ 3.1 has also been made easier with a result comparison tool that can also be used to verify initial and confirmatory HLA typing data.
Figure 3: TypeStream Visual™ provides the user with a streamlined analysis experience.

NGS is complicated – but your workflow doesn’t have to be. Get in touch to learn how you can adopt the AllType™ FASTplex™ NGS method for HLA typing that’s simple, fast and best-in-class.
Enquire about this article
"*" indicates required fields
