
PCR-SSP for HLA genotyping (using Sequence-Specific Primers) was initially described in 1988 and exploded in popularity in the early 1990s with publications defining the potential for use in solid organ transplantation. The commercialisation of PCR-SSP led to widespread adoption of this method, which remained largely unchanged for a number of decades. PCR-SSP overcame the obstacles presented by serological HLA typing: limited, polyclonal antisera, the need to live cells, and the inability to ‘capture’ raw data for future reference. However, PCR-SSP requires gel electrophoresis for the separation of amplified DNA fragments, and ethidium bromide (a suspected carcinogen) and UV transillumination present significant health and safety concerns.
LinkSēq™
Launched in 2010, the LinkSēq™ HLA product range is a ground-up rethink of the PCR-SSP system. LinkSēq™ 1 utilises qPCR instrumentation and SYBR Green chemistry as opposed to gel electrophoresis to capture the assay end point. PCR amplification occurs if sequence-specific primers bind to complimentary sequence in the DNA added. SYBR Green is inherent in the LinkSēq™ [PCR] Buffer and preferentially binds double-stranded DNA to emit green fluorescence; fluorescence increases as the number of amplicons increase. After thermal cycling, PCR amplicons are heated across a temperature range using a qPCR instrument. The resultant amplicon will ‘melt’ (become single-stranded) at a specific temperature, depending on e.g. the size, G:C content and sequence motifs captured within the DNA fragment. When each amplicon reaches its specific melting temperature SYBR Green fluorescence decreases rapidly; this is captured by the qPCR instrument.
Speed and Simplicity
LinkSēq™’s use of qPCR instrumentation and SYBR Green chemistry offers speed and simplicity. After PCR set-up (10 minutes) and thermal cycling (60 minutes), melt-curve analysis takes only 10 minutes. Results are exported from the qPCR instrument in a matter of seconds and analysed using One Lambda’s SureTyper™ analysis software. Analysis is largely automated (whilst allowing the user to interrogate raw data) such that an HLA type is available for reporting within minutes. Overall, LinkSēq™ reduces the time from DNA to results to only 90 minutes (Figure 1). The workflow is simple: after set-up the plate is sealed and both thermal cycling and melt-curve analysis are performed on the qPCR instrument. After data review the tray may be thrown away without the need for further processing.

Figure 1: LinkSēq™ workflow.
Flexibility
LinkSēq™ also offers a flexible solution. Primer sets are available in a variety of different formats: 384-Well and 96-Well trays, and 8-Well strips (Figure 2). For example, the popular 1575C product offers 11-loci HLA genotyping (HLA-A, -B, -C, -DRB1, -DRB3/4/5, -DQA1, -DQB1, -DPA1 and -DPB1) from a single 384-Well tray. Alternatively, the 3030C kit allows the HLA-A, -B, -DRB1 and -DRB3/4/5 genotyping of two samples using a single 96-Well tray. Single-Locus kits are available in 384-Well and 96-Well formats, allowing the user to test for selected loci or perform repeat typing. For specific applications requiring fewer primer sets (e.g. HLA-B27 disease association and HLA-B*57:01 drug hypersensitivity genotyping), three samples can be tested using a single 8-Well strip. LinkSēq™ also offers ancillary products such as the LinkSēq™ Wipe Test to monitor for contamination in the PCR environment.
Figure 2: LinkSēq™ 384-Well format.
The LinkSēq™ range has also been validated across a number of qPCR instruments (for example, models from the Life Technologies QuantStudio and Roche LightCycler ranges) with guidance available for how to set-up the necessary protocols for thermal cycling/melt-curve and data retrieval. Importantly, although thermal cycling and melt-curve analysis are typically performed on the qPCR instrument, these processes can be uncoupled. This flexibility means LinkSēq™ genotyping is easily scalable, enabling laboratory throughput. For example, using six thermal cyclers and one qPCR instrument, six samples could be HLA typed for all 11-loci in approximately three hours: PCR set-up (10 minutes/sample), simultaneous thermal cycling (60 minutes), and back-to-back melt-curve analysis (10 minutes/sample).
Innovative Product Portfolio
The LinkSēq™ product portfolio is continually improving and expanding with novel and innovative products. An alternative to the 1575C product for 11-loci HLA genotyping is the 1580C Single Antigen Bead Resolution ‘SABR’ product. This 384-Well kit includes additional primer sets such that key alleles can be distinguished within serological groups (e.g. HLA-DQ6; Figure 3). At launch, sufficient primers were incorporated to provide resolution that complements solid-phase antibody specification assays provided by multiple manufacturers. More recently One Lambda have introduced Multi-Target Assays to increase the resolution of LinkSēq™ further still: multiple HLA-specific primer sets occupy the same well, allowing for more than 384 primer pairs in the 384-Well SABR product. The LinkSēq™ product range also extends beyond HLA genotyping. The LinkSēq™ KIR assay types all 15 Killer Immunoglobulin-like Receptor genes and two pseudogenes, including both full length and deleted forms of 3DP1 and 2DS4. In the transfusion arena, LinkSēq™ HPA typing kits provide Human Platelet Antigen typing for HPA-1 through HPA-11, and HPA-15. LinkSēq™ ABO and LinkSēq™ RBC assays provide genotyping for assays typically reliant on antisera and for which discordant serological reactions may occur (e.g. for weakly expressed A antigen). LinkSēq™ ABO 2 (for ABO blood grouping) separates A1 from non-A1, B from B3, Bweak and Bel etc., whilst the LinkSēq™ RBC assay targets RhD, RhCE, MNS, Kell, Duffy, Kidd, Lutheran, Diego, Cartwright, Scianna, Dombrock, Colton, Landsteiner-Wiener and Haemoglobin.
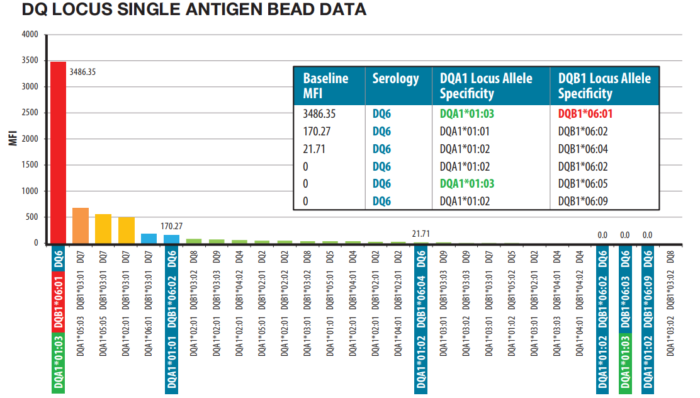
Figure 3: LinkSēq™ SABR aids patient-donor compatibility assessment. Class II DQ data showing strong positive reactivity for allele specific DQB1*06:01 while other DQ6 allele specific antibodies are not present. DQ6 alleles are outlined in blue. DQA1*01:03 is outlined in green. DQB1*06:01 is outlined in red and is the only positive DQ6 allele.
Powerful Software
One Lambda’s SureTyper™ software offers an intuitive user experience when analysing LinkSēq™ 3 4. Once qPCR data are imported, genotyping results are generated automatically. Should the user wish to look behind the result and interrogate raw data, e.g. in the event of a less common allele call, SureTyper™ expands into four quadrants with melt-curve data transformed into peaks analogous to bands in a PCR-SSP electrophoresis gel (Figure 4). The user can ‘flag’ questionable assays and perform a What-If analysis whereby SureTyper™ considers the result as if it were positive, and as if it were negative. SureTyper™ also utilises robust points of reference. For example, SureTyper™ for ABO uses ABO allele frequency information from the Blood Group Antigen Facts Book and the Erythrogene search engine for blood group genes 5 6 whilst SureTyper™ for HLA utilizes the common and well-documented HLA alleles catalogue by Mack et al. (2013) 7. This enhances confidence when assigning common alleles and encourages the user to further evaluate data when ‘rare’ alleles are present.
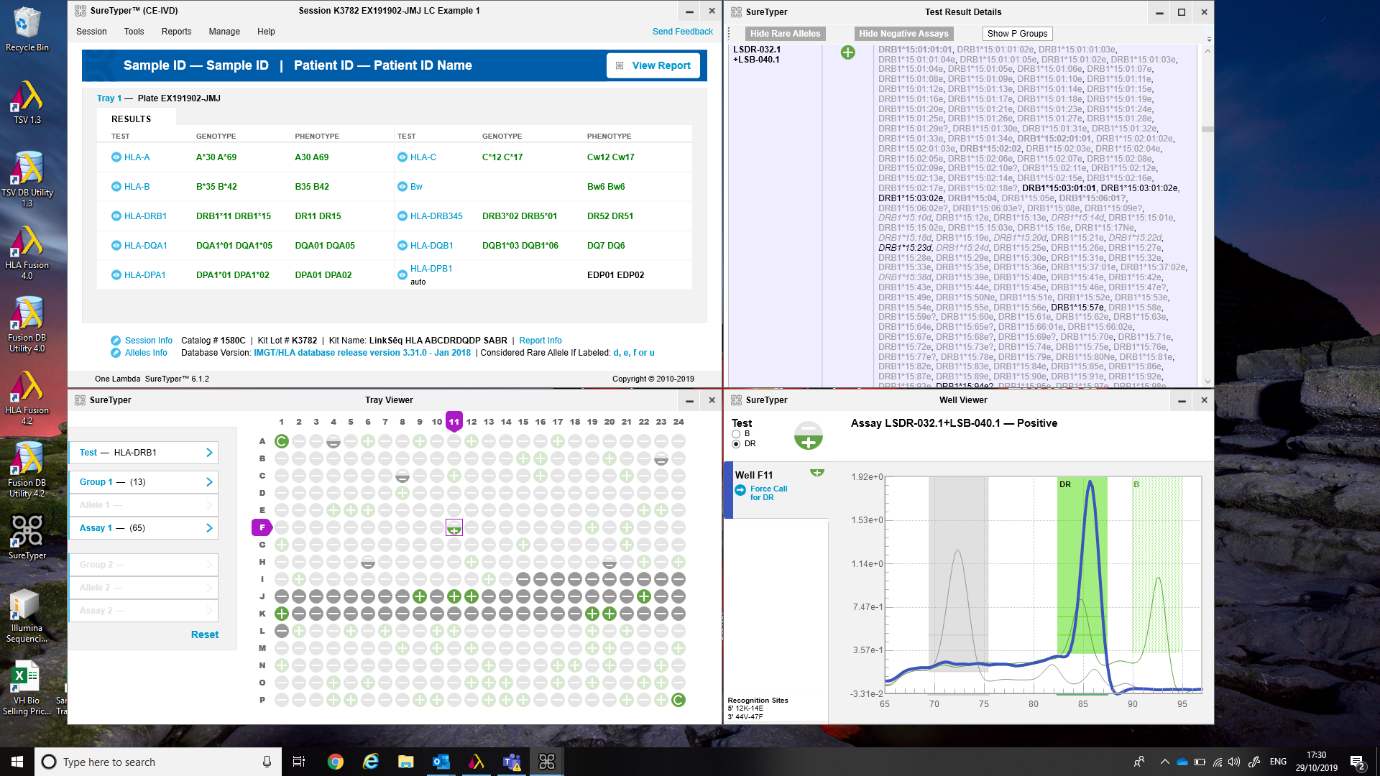
Figure 4: SureTyper™ for HLA.
Quality-Assured
Many LinkSēq™ HLA products are CE-IVD, as is the SureTyper™ for HLA software. In addition to the availability of a LinkSēq™ Wipe Test for contamination monitoring, assay quality is ensured by the presence of controls integral to the LinkSēq™ assay: a No Template Control flags contamination; a DNA Quality Control indicates where fragmented DNA may compromise result quality, and an Internal Control is present in each well to confirm the authenticity of a negative result. SureTyper™ software also includes check-points to ensure sample and donors identifiers are associated with the correct data set.
One Lambda’s LinkSēq™ range and SureTyper™ analysis software are easily integrated into your laboratory and provide a fast, flexible solution for the genotyping of HLA, KIR, HPA, ABO and red blood cell antigens.
References
- One Lambda LinkSēq™ Typing Kits. https://www.onelambda.com/en/products-services/products/molecular-typing/LinkSēq™.html
- ABO Blood Subgrouping and Implications in Transplantation. https://www.youtube.com/watch?v=rGrWCiGHujs&feature=youtu.be
- One Lambda SureTyper™ Software https://www.onelambda.com/en/product/SureTyper™-software.html
- How To: Analyze LinkSēq™ HLA Typing Kits using SureTyper™ Software v6.0. https://www.youtube.com/watch?v=R9mjRLh9jzE&feature=youtu.be
- Marion E. Reid, M.E, Lomas-Francis, C. and Olsson, M.L. Blood Group Antigen Facts Book, third edition. ISBN 978-0-12-415849-8.
- Möller, M., Jöud, M., Storry, J. R., & Olsson, M. L. Erythrogene: a database for in-depth analysis of the extensive variation in 36 blood group systems in the 1000 Genomes Project. http://www.erythrogene.com, v0.7 (22 Dec 2016). Blood Advances, 1(3): 240-249. https://doi.org/10.1182/bloodadvances.2016001867
- Mack, S. et al. Common and well-documented HLA alleles: 2012 update to the CWD catalogue. Tissue Antigens, 81:194-203.
Enquire about this article
"*" indicates required fields