
Extracting reliable DNA from your samples is vital, but with so many extraction methods, how can you decide which is best for your intended results?
Read our guide to DNA extraction methods to find the right solution for your lab:
The extraction of genomic DNA is a crucial first step in many different types of studies. The reliability of your DNA extraction method plays a huge role in providing you with accurate and meaningful downstream results. Efficient DNA extraction leads to accurate and meaningful results at your end point, whether that be gel electrophoresis, PCR based assays or metagenomics. Not having a good DNA extraction method can result in the presence of PCR inhibitors, a reduced yield and loss of DNA quality. Other aspects to consider are your intended processing time, the scalability of your method, and cost.
What are the different types of DNA extraction?
There are a few different ways you can extract DNA from your sample:
- ethanol precipitation
- magnetic bead extraction
- and column-based methods
There are different features and benefits of each method, and these can help you decide which one is the best for you. All methods involve cell lysis followed by separation and purification, but use different principles (precipitation, adsorption to beads, or adsorption to columns).
So, which DNA extraction is best for your workflow?
1. Ethanol precipitation
Ethanol precipitation is a classic method that allows you to extract your DNA using detergents to release DNA. This occurs through a multistep centrifugation process using phenol, chloroform and isoamyl alcohol. After these centrifugation steps, the DNA remains in the aqueous layer while the proteins and lipids move into the organic layer. The aqueous layer is removed by careful pipetting and layered onto the next chemical for centrifugation. The result is a pellet which would contain your purified DNA which can be resuspended in buffer or water.
Pro: If carried out carefully, ethanol precipitation can give high yield and good quality DNA at relatively low reagent cost.
Con: However, it uses hazardous chemicals and is time intensive. It is not commonly automated in modern labs, where column or bead methods are preferred for scalability and convenience.
2. Magnetic bead-based methods
Magnetic bead-based DNA extraction methods use magnetic beads to isolate DNA. After cell lysis, the DNA within a sample is captured by the magnetic beads which are coated in a reagent which allows for the binding of the DNA. When a magnetic field is applied, this separates the DNA-bound beads from the remaining contaminants allowing them to be washed away. The DNA can then be eluted through the addition of a buffer to release the DNA from the beads.
Pro: These methods are particularly well suited to automation, reducing hands-on time and contamination risk. They’re widely used in clinical workflows because sealed, pre-packaged kits enhance safety.
Con: While automation can improve cost-effectiveness at scale, the consumables are not necessarily cheaper than manual columns.
Note: magnetic bead extraction is not the same as bead beating, which is a physical tissue disruption step sometimes used prior to extraction.
3. Column-based methods
Column-based methods utilise a purification matrix to bind DNA. This method uses a chaotropic salt and a low pH buffer to allow the DNA to bind to matrix, while other components flow through it and are washed away. The sample is pushed through the column by a series of centrifugation or vacuum steps, and the purified DNA is eventually released and eluted in the same way.
Pro: Manual spin columns are cost-effective and widely used.
Con: Scaling up manually can be labour-intensive. However, automated platforms are available for higher throughput.
Magnetic bead vs column-based DNA extraction solutions
Here, we compare examples of a magnetic bead-based method from Precision System Science (PSS) and column-based method from Molzym, both available from VH Bio.
These methods can offer a safer, faster, more convenient alternative to ethanol precipitation and come with technical support from VH Bio to help you through the process.
The magnetic bead-based solutions from PSS
The magLEAD 12gC Link by PSS is a platform supplied by VH Bio which performs magnetic bead-based DNA extraction. The instrument has less than 5 minutes of hands-on time and can extract the DNA from various sample types including whole blood, plasma, serum, swabs, sputum, stool, saliva, cerebrospinal fluid, urine, formalin-fixed paraffin embedded tissues and many more.
With automation, there is a reduction in contamination risk as the instrument uses only pre-packaged reagents and has an inbuilt UV lamp for decontamination. The magLEAD 12gC Link allows you to process up to 12 samples in as little as 26 minutes (sample type dependant) in a CE-IVDR certified workflow which integrates into laboratory information systems. Extraction reagents and plasticware can be stored at room temperature, omitting the need for fridge space which can be beneficial for storage space. Using this instrument, you can extract total nucleic acid and/or total RNA for your downstream applications as shown in Figure 1. While not “low cost” in absolute terms, it is cost-effective compared with larger automation platforms.
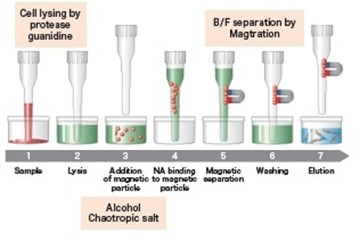
Figure 1: The Magtration® technology workflow using the MagDEA Dx MV nucleic acid extraction kits. 1). Sample is added, 2). Proteins are lysed within the sample through chaotropic agents. 3). Magnetic beads are added. 4). The phosphate group of nucleic acid binds to a magnetic bead carrier by a complex mechanism 5). The magnetic particles are separated from other components within the solution 6). The magnetic beads are washed removing other debris. 7). Nucleic acids are eluted from the magnetic beads.
Read more: Introducing the MagLEAD 12gC Link: Precision Nucleic Acid Extraction for Your Lab
The column-based method from Molzym
Molzym have developed the MolYsis technology, a column-based DNA extraction with the added benefit of depleting host DNA presence (Figure 2).
The MolYsis technology has been developed to provide laboratories with the ability to increase the efficiency and accuracy of microbial DNA analysis, allowing users to identify bacteria and fungi which otherwise may have been lost. This is achieved through the removal of human ‘host’ DNA, which in the case of microbiology, is background noise which can mask the desired DNA.
This method is ideally suited for culture-independent molecular analysis within the clinic, PCR & NGS applications and for low load samples, and has a shorter turn-around time to result. This method can be performed manually or using the SelectNA™plus automated instrument. The appropriate type of kit is based on your desired sample type and downstream application as shown in Figure 3.
The SelectNA™plus instrument allows users to process 1-12 samples in around 1.5 to 2hrs after loading the instrument by utilising an innovative vacuum-controlled technology to move the sample through the column. The MolYsis kits are also equipped with DNA free plasticware which ensures users that their resulting DNA eluate is contaminant free. These kits have been validated with a wide range of sample types including body fluids, swabs and tissues.
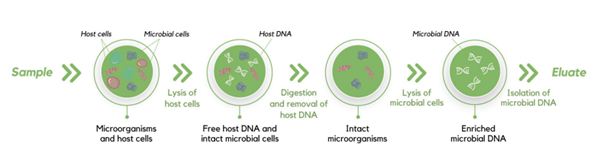
Figure 2: The MolYsis technology. The column-based DNA extraction process from Molzym includes host DNA depletion. The MolYsis technology begins with the lysis of the host cells which exposes the host DNA for digestion by DNases. The debris is then moved through the column and washed away. The remaining microbial cells are then lysed separately, exposing the microbial DNA which will then be captured by the column matrix. This can then be eluted after the remaining debris is washed away.
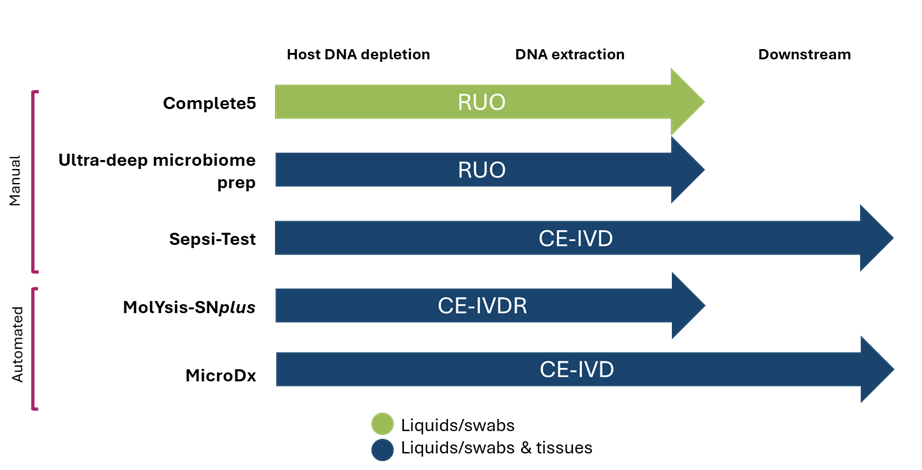
Figure 3: The different kits available for column-based DNA extraction from Molzym.
The manual kits available for host DNA depletion and DNA extraction include the Complete5 kit which is research use only (RUO) and is appropriate for liquid samples and swabs. The Ultra-Deep Microbiome Prep kit is RUO, offers the same workflow and is compatible with liquid, swab and tissue samples. All kits with a blue arrow include a pre-digestion step for viscous or tissue samples and the DNA free plasticware to perform this step. The SepsiTest-UMD kit is CE-IVD certified and comes with 16S/18S PCR reagents and Sanger sequencing primers for a full sample to identification workflow. The automated kits are both accredited workflows. The MolYsis-SNplus™ IVD allows the end user to deplete host DNA and extract microbial DNA to be used with a downstream method of the user’s choice such as metagenomics. The Micro-DX™ kit allows the end user to deplete host DNA, extract microbial DNA and identify the extracted microorganisms with included 16S/18S PCR reagents and Sanger sequencing primers.
How to choose your DNA extraction method?
There are many benefits to both the magnetic bead-based and column-based methods, summarised in Figure 4. Deciding on what method is the most appropriate for your workflow will ultimately depend on your:
- Sample type (Are the tissue samples in need of pre-processing first? Are they fresh/frozen samples? Are they hazardous?)
- Lab space and available equipment (Do you require a manual extraction or automation?)
- Budget
- Sample numbers
- Downstream applications (PCR, microbial metagenomics)
- Project focus (What do you want to detect in your sample? i.e. human DNA or microbial DNA or all nucleic acid contents?)
In conclusion, there are benefits to all DNA extraction methods, but particular approaches can enhance your final outcome based on your overall needs.
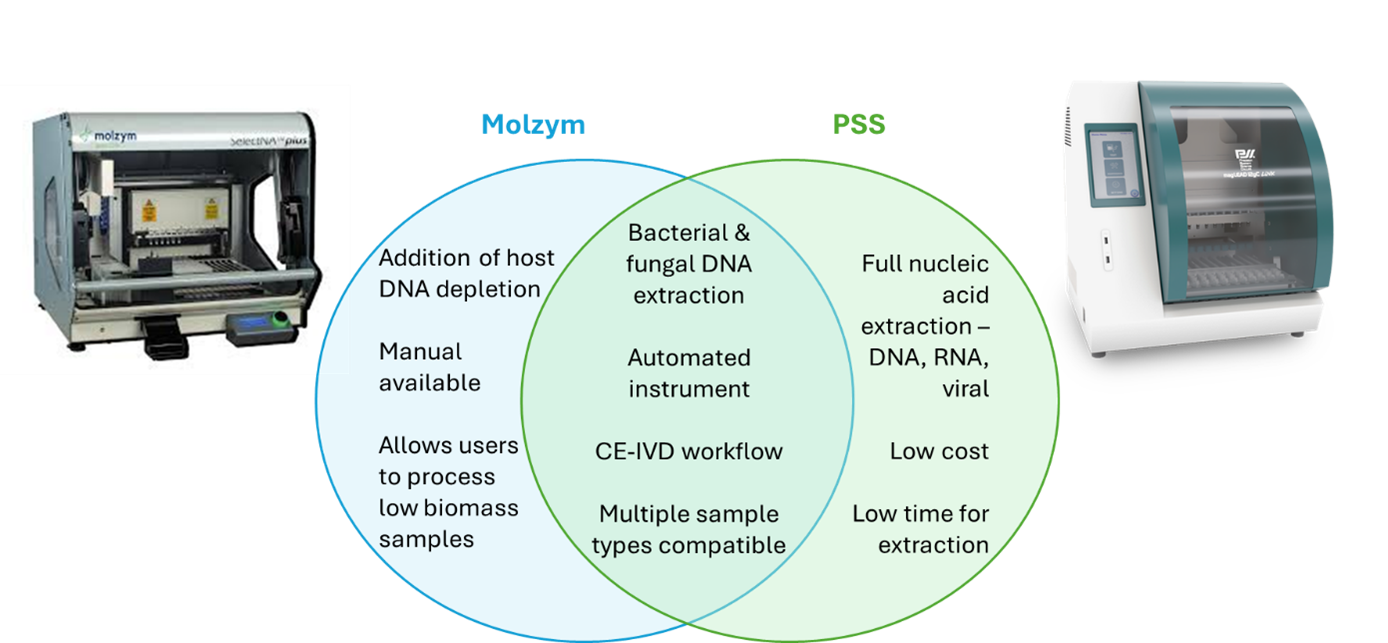
Figure 4: Summary of the MolYsis and Magtration technologies from Molzym and PSS, respectively.
Conclusion
The Molzym solutions allow their users to utilise host DNA depletion and are available in manual and automated formats with the result being extracted microbial DNA. The MolYsis technology allows users to process low biomass samples for downstream microbial identification through a column-based method.
The PSS instrument enables full nucleic acid extraction, allowing users to view the full nucleic population within their sample. This extraction method is a magnetic bead-based method and is a low cost compared other automation platforms for extraction.
Both methods are compatible with multiple sample types in a CE-IVDR certified workflow.
Has this guide helped you determine the right method for your DNA extraction? Request an order through our website or get in touch with our team who can answer your questions to help you achieve accurate and meaningful downstream results from DNA extraction.
Enquire about this article
"*" indicates required fields
