Background
It has been over 60 years since the first successful kidney transplant was performed and transplantation remains the single most effective treatment for patients suffering with end stage renal disease (ESRD); however, demand outweighs supply by more than 2:1 (Figure 1).
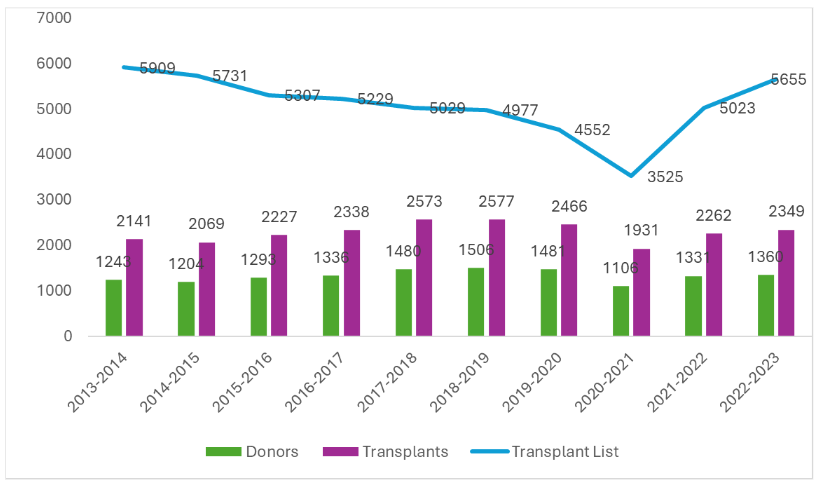
FIGURE 1: Deceased donor kidney programme in the UK 01/04/2013 – 31/03/2023 (1).
Advances in immunosuppression, immunotherapy, surgical procedures and histocompatibility and immunogenetics (H&I) testing in those 60 years have seen the rate of graft survival improve significantly. Recent NHSBT data demonstrate that, while the UK has some of the best graft and patient survival rates in the world (2), graft and patient survival has plateaued over the past 20 years (3). Furthermore, a study reviewing 22,000 kidneys transplanted in the UK between 2000 and 2013 found a staggering 20% of transplanted kidneys failed in the first five years, with alloimmune pathology a major cause of allograft loss (25%).
Post-Transplant Monitoring
In the UK, post-transplant monitoring involves reviewing levels of biochemical biomarkers in recipient blood and urine. For example, serum creatinine, urea and electrolytes (used to estimate glomerular filtration rate; eGFR); tests that are both inexpensive and available 24/7 at most hospital laboratories. Immunosuppressants such as tacrolimus and sirolimus are also monitored to identify whether dosage is too low or too high and, in some cases, confirm non-adherence at time points when markers of rejection are elevated. Immunosuppressants can also exhibit cytotoxic effects, so liver enzymes to assess liver function may also be monitored.
Despite Luminex bead-based methods for detecting donor-specific antibodies (DSA) being more sensitive and specific than serum creatinine levels for detecting antibody-mediated rejection (ABMR), few H&I laboratories routinely monitor for de novo DSA post-transplant. In many laboratories, post-transplant DSA monitoring is triggered following an observed rise in creatinine or due to clinical symptoms. If DSA are identified, a patient may require additional therapy to minimise rejection and a biopsy may be indicated to confirm whether allograft injury is due to infection or is immunologically driven (either cellular- or antibody-mediated rejection).
Herath and colleagues (5) argue that current methodologies for post-transplant monitoring detect allograft injury too late for available treatments to be efficacious and that non-invasive methods that can detect allograft dysfunction much earlier are required (Figure 2).
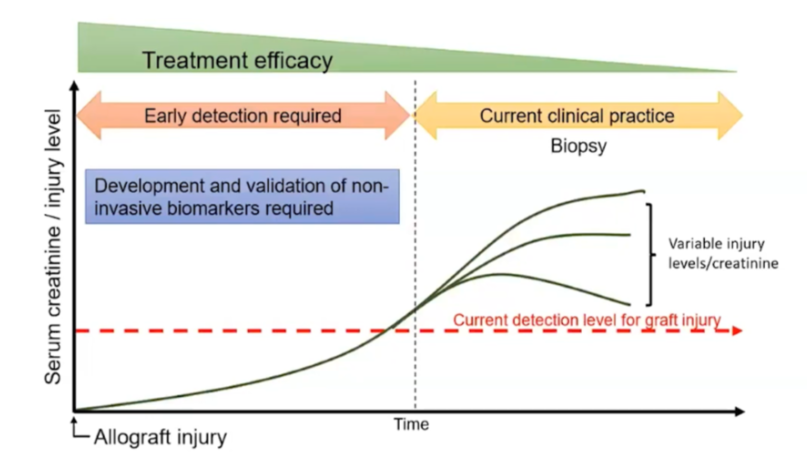
FIGURE 2: Herath et al. argue that biopsy is usually late when triggered by a rise in creatinine and express the need for a timely, objective, and non-invasive diagnostic test (5).
What is dd-cfDNA?
Donor-derived cell-free DNA (dd-cfDNA) is naturally shed by donor graft cells undergoing necrosis and apoptosis and can be isolated from human plasma, quantified, and monitored using cell-free DNA isolation techniques and next-generation sequencing (NGS) technology; effectively providing the opportunity to assess allograft health via a method that is minimally invasive.
A review by Oellerich et al. (6) described the need to improve personalised immunosuppression to reduce premature graft loss and identified dd-cfDNA as having demonstrable application in the following areas.
• Detection or exclusion of graft injury or rejection;
• Earlier detection of transplant injury;
• Earlier diagnosis of subclinical ABMR;
• Earlier intervention;
• Alternative to biopsy;
• Indicator of under-immunosuppression;
• Personalised immunosuppression;
• Evaluation of infectious complications on allografts.
Even a minor elevation in %dd-cfDNA – anywhere between 0.2% and 1% – has been found to correspond with graft dysfunction (7,8,9). Through a prospective, multi-centre study involving 1,092 transplanted kidneys, Bu and colleagues established that a 0.5% rise in %dd-cfDNA was an earlier marker of allograft dysfunction than creatinine and DSA, with one case showing a rise in %dd-cfDNA 91 days before a rise in DSA, as well as predicting a 25% fall in eGFR over three years. Patients with a persistent increase in %dd-cfDNA had a poorer prognosis.
Conversely, where dd-cfDNA monitoring demonstrated immune quiescence, this negated the need for monitoring biopsies (7). Reducing monitoring biopsies may be especially impactful for cardiothoracic teams wishing to reduce the number of endomyocardial biopsies in heart transplant recipients. Feingold et al. (10) reported an 81.8% reduction in endomyocardial biopsies in the first 34.5 months after adopting dd-cfDNA testing, with early results showing no significant difference in clinical outcome between recipients who were being biopsied versus those being monitored using dd-cfDNA testing. Fewer biopsies would lead to a significantly better patient experience post-transplantation.
Consensus agreement in the clinical utility of dd-cfDNA monitoring
The American Society for Transplant Surgeons (ASTS) released a position statement in July 2023, summarising:
“Collectively, these and other data point to significant clinical utility of dd-cfDNA in clinical decision formation in kidney transplant recipients, with low dd-cfDNA levels providing evidence of immune quiescence and providing the ability to forego unnecessary biopsies, moderately elevated or rising levels signalling a markedly elevated risk of rejection, and higher levels providing a high degree of confidence in the presence of underlying antibody-meditated rejection (ABMR). The demonstrated clinical utility of dd-cfDNA includes a significant improvement relative to serum creatinine measurement in the detection of both [T-cell mediated rejection] TCMR and ABMR. As a marker of allograft injury, dd-cfDNA is an earlier indicator of allograft injury than de novo DSA and of serum creatinine” (11).
Additionally, the European Solid Organ Transplant (ESOT) consensus conference in November 2022 reached 100% agreement in recommending:
“that clinicians consider measuring serial plasma dd-cfDNA testing in patients with stable graft function and with acute graft dysfunction to exclude the presence of subclinical antibody-mediated rejection” when asked “is dd-cfDNA a reliable diagnostic tool for acute rejection monitoring compared with standard of care (eGFR/creatinine monitoring or for cause biopsy)?” (12).
With over 30 published studies demonstrating the clinical utility of dd-cfDNA monitoring, it seems prudent to explore this tool for potential early intervention and avoidance of unnecessary biopsies.
How does dd-cfDNA monitoring work?
Devyser, a Swedish company specialising in developing pioneering and highly efficient diagnostic genetic tests, has developed a simple and robust NGS-based method for detecting dd-cfDNA for post-transplant monitoring (13). In July 2023, Devyser partnered with One Lambda, Inc., completing One Lambda’s extensive transplant diagnostics portfolio with NGS-based post-transplant monitoring products, including ‘Accept cfDNA’ for solid organ transplantation and ‘Chimerism’ for haematopoietic progenitor cell transplantation (HPCT).
Accept cfDNA is CE-IVD certified for diagnostic use in the European and UK markets and, designed with IVDR regulations in mind, is the first dd-cfDNA monitoring method to be IVDR-certified too. The assay has been validated using both EDTA and STRECK tubes, both automated and manual cfDNA isolation methods across a variety of platforms (by QIAGEN, Promega and ThermoFisher), and across multiple Illumina NGS platforms (MiSeq, iSeq and MiniSeq models). The NGS method (Figure 3) has a streamlined workflow, involving two relatively short PCRs to amplify 50 population-independent insertion/deletions (indels), before adding library indexes to amplicons (barcoding).
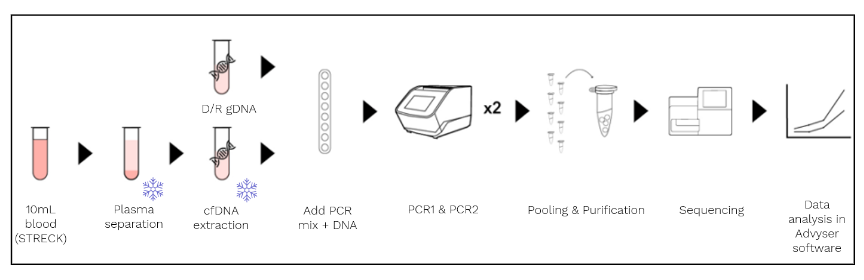
FIGURE 3. The One Lambda Devyser Accept cfDNA screening (above) and monitoring (below) workflows. Key: * = stages at which samples can be frozen pre-PCR.
After barcoding, samples can be pooled in a single-tube for amplicon size-selection and purification of the library pool, eliminating the need for several reagent and amplicon transfer steps. By targeting indels, small amplicons (less than 75 base pairs in length) can be sequenced using faster sequencing programmes than those targeting full gene lengths, and by carefully selecting population-independent indels, fewer targets are required, minimising reagent and flow cell usage. Devyser have conveniently aligned their Accept cfDNA protocol with their post-HPCT Chimerism protocol, meaning that final libraries for both assays can be pooled and run on the same flow cell; Devyser even provide a calculator tool to maximise flow cell usage and minimise waste.
Once sequencing is complete, outputted data is imported into the assay’s dedicated Advyser Solid Organ software. The software reviews quality metrics as it rapidly analyses results for all 50 indels, presenting the user with a list of informative markers, indicating where genetic differences are identified between the donor and recipient. The software calculates %dd-cfDNA relative to patient cfDNA and presents this to the user, graphically, for longitudinal monitoring. Graphs and results can then be exported to Laboratory Information Management Systems (LIMS). Where a previous donor’s DNA is available, the software can also monitor dd-cfDNA from two different donors alongside each other.
Summary
The necessity for a non-invasive, timely identifier of allograft injury and for personalised immunosuppression has been established to improve graft outcomes and patient survival (4,5,6). dd-cfDNA monitoring has demonstrable clinical utility, with consensus and evidence-based agreement for its application among the ASTS and ESOT panel of experts (11,12). Accept cfDNA is a non-invasive, robust, and simple NGS solution for post-transplant monitoring of transplanted kidneys (with clinical studies in progress for other solid organs). By targeting indels over single nucleotide polymorphisms (SNPs), Accept cfDNA reduces sequence-related artefacts by x100 and can detect dd-cfDNA as low as 0.1%, giving this assay the greatest sensitivity on the market. With just 45 minutes of hands-on time, Devyser One Lambda Accept cfDNA not only halves the time required by technical staff compared to a competitor’s product (AlloSure, CareDx) but also brings unrivalled sensitivity, quality, and expertise in %dd-cfDNA monitoring to the H&I laboratory, whist also reducing the redundancy time of Illumina instrumentation. Most importantly, earlier detection of allograft dysfunction should enable earlier intervention, improve allograft outcomes and improve patient outcomes.
Want to find out more?
Get in touch with your transplant diagnostic colleagues at V.H. Bio Limited to discuss how to bring Accept cfDNA into your testing repertoire, improving transplant outcomes and the post-transplant experience of patients.
References
1. Figure 1: https://nhsbtdbe.blob.core.windows.net/umbraco-assets-corp/30139/section-5-kidney-activity.pdf [accessed 20/12/23]
2. Graham et al. (2022) Mean lifetime survival estimates following solid organ transplantation in the US and UK, Journal of Medical Economics, 25:1, 230-237
3. Graft and patient survival: https://nhsbtdbe.blob.core.windows.net/umbraco-assets-corp/30188/activity-report-2022-2023-final.pdf [accessed 21/12/23]
4. Burton et al. (2019) Causes of renal allograft failure in the UK: trends in UK Renal Registry and National Health Service Blood and Transplant data from 2000 to 2013. Nephrol Dial Transplant. 34(2):355-364
5. Herath, S. et al. (2019) Advances in Detection of Kidney Transplant Injury. Mol Diagn Ther 23: 333–351
6. Oellerich et al. (2019). Absolute quantification of donor-derived cell-free DNA as a marker of rejection and graft injury in kidney transplantation: Results from a prospective observational study. Am. J. Transpl. 19, 3087–3099.
7. Bu et al. (2022) Clinical outcomes from the Assessing Donor-derived cell-free DNA Monitoring Insights of kidney Allografts with Longitudinal surveillance (ADMIRAL) study, Kidney International 101:793-803
8. Halloran et al. (2022) The Trifecta Study: Comparing Plasma Levels of Donor-derived Cell-free DNA with Molecular Phenotype of Kidney Transplant Biopsies, J Am Soc Nephrol 33: 387-400
9. Bloom et al. (2017) Circulating Donor-Derived Cell-Free DNA in Blood for Diagnosing Active Rejection in Kidney Transplant Recipients (DART) Study Investigators. Cell-Free DNA and Active Rejection in Kidney Allografts. J Am Soc Nephrol. 28(7):2221-2232
10. Feingold et al. (2023) Short-term clinical outcomes and predicted cost savings of dd-cfDNA-led surveillance after pediatric heart transplantation. Clin Transplant. 37:e14933
11. ASTS Position Statement: https://asts.org/docs/default-source/position-statements/dd-cfdna-position-statement.pdf [accessed 23/09/23]
12. ESOT Consensus Conference Highlights: https://esot.org/wp-content/uploads/2023/03/EM012464_TLJ_3_0_Scientific_Highlights_Report_230316_V0-7_SMB.pdf p.22-25 [accessed 23/09/23]
13. Pettersson et al. (2024) Development and performance of a next generation sequencing (NGS) assay for monitoring of dd-cfDNA post solid organ transplantation Clin Chim Acta 552:117647
14. Schirmer et al. (2015) Insight into biases and sequencing errors for amplicon sequencing with the Illumina MiSeq platform, Nucleic Acids Research 43(6):e37
Enquire about this article
"*" indicates required fields
