Introduction
Human neutrophil antigens (HNA) are a family of epitopes expressed on the glycoproteins of granulocytes, 99% of which are neutrophils; hence the terms ‘granulocytes’ and ‘neutrophils’ being used interchangeably in the literature. There are five known HNA systems to which these epitopes belong, encoded by alleles that are differentiated by single nucleotide polymorphisms (SNPs) in known coding positions along the glycoprotein genes (Table 1).
Neutrophils play a critical role in the immune response to pathogens: acting as the first line of defence in innate immunity, travelling to the source of the infection to initiate an inflammatory response, recruiting and upregulating the production of enzymes, peptides, reactive oxygen species, and inflammatory cytokines (e.g. TNF-α) to clear pathogens by means such as apoptosis and phagocytosis.
Neutrophils express Fc gamma receptors (FcγRs) which interact with the invariable (Fc) portion of IgG molecules. FcγrIIIb (also known as Cluster of Differentiation 16 or CD16) expression depends on the stage of neutrophil development, with high expression seen on mature neutrophils. CD16 activates Natural Killer cells and can moderate both T cell and antibody-dependent cytotoxicity.
Due to the integral role played by neutrophils in host defence, neutropenia – a decline in circulating neutrophils – can result in recurrent and potentially fatal infections. Neutropenia may be mild, moderate or severe; idiopathic; or caused by infections, haematological malignancies (e.g., leukaemia), treatment (e.g. radiation therapy), nutritional deficiencies, or anti-HNA antibodies.
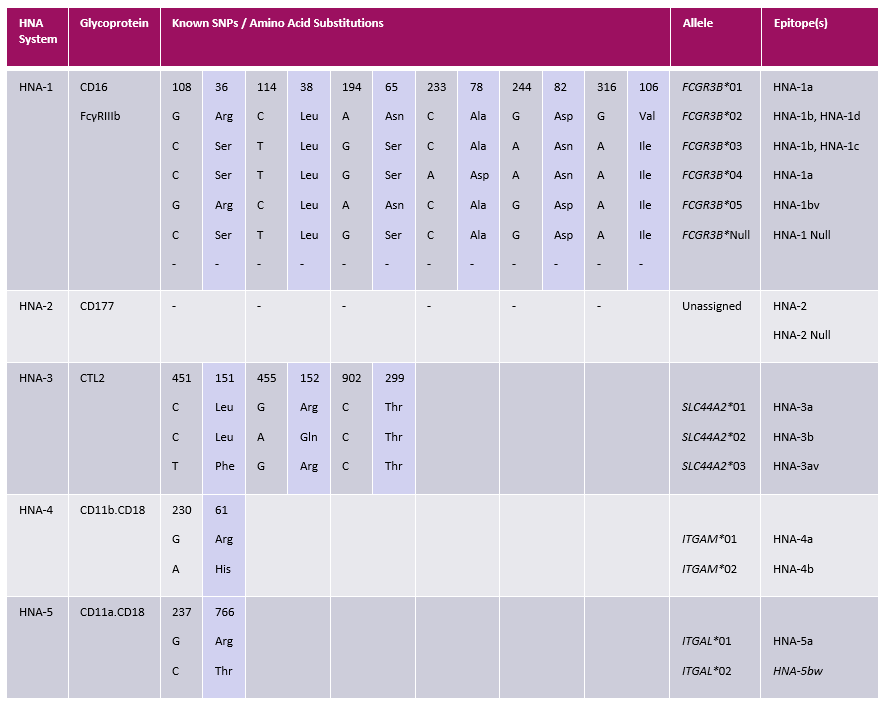
Table 1. Known polymorphic regions of genes encoding HNA epitopes. Adapted from Flesch et al. (2018) and Browne et al. (2021).
Clinical Implications of Anti-HNA Antibodies
Neutropenia
The generation of anti-HNA antibodies may be autoimmune or alloimmune in nature. Autoimmune neutropenia of infancy (ANI) can be primary or idiopathic, affecting newborns and infants with no known cause, often resolving itself within the first few years. ANI can also be secondary to autoimmune conditions (e.g. Crohn’s Disease), predominately affecting older children and adults over many years.
Alloimmune neutropenia (AIN) occurs when a patient has developed antibodies against non-self HNA. In foetal/neonatal alloimmune neutropenia (FNAIN), maternal antibodies directed against paternal HNA expressed on foetal granulocytes can cross the placenta, destroying foetal neutrophils, leaving the foetus/neonate vulnerable to potentially life-threatening infections. FNAIN usually resolves within the first few months as the maternal antibodies are cleared from circulation while the immune system of the infant develops.
Transfusion Reactions
Anti-HNA antibodies can lead to adverse reactions following the transfusion of blood products. Febrile non-haemolytic transfusion reactions (FNHTR) impact between 1-3% of transfusions and are characterised by an increase in temperature by greater than 1°C, chills and/or hypertension. This has been attributed to either bacterial contamination or antibodies against HLA Class I, HLA Class II, or HNA.
When antibodies against HNA are present in the plasma being transfused, they can target HNA on recipient granulocytes causing absolute neutropenia; a conditioned called transfusion-related alloimmune neutropenia (TRAIN). Thankfully, the incidence of TRAIN is rare.
Transfusion-related acute lung injury (TRALI) is a potentially life-threatening condition, with symptoms such as pyrexia, hypoxia, dyspnoea and pulmonary oedema manifesting within six hours of receiving a transfusion. Antibodies against HLA Class I, HLA Class II and HNA-1, -2, and -3a have been implicated in TRALI cases, though it can ensue from fluid overload in patients with pre-existing comorbidities without antibody involvement. Cases are considerably lower since TRALI reduction measures were introduced, screening donor plasma for antibodies and exclusively recruiting male donors for the avoidance of TRALI.
Transplantation Rejection
Anti-HNA antibodies have been associated with haematopoietic stem cell transplant rejection. Incompatibility can result in neutropenia, delayed engraftment, non-engraftment, and rejection (Alswied et al., 2024).
There have also been several case reports in the UK and globally of HNA-3a antibody involvement in kidney transplant rejection (Key et al., 2019; Cannon et al., 2023), where a positive flow cytometry crossmatch was observed in the absence of HLA antibodies detectable by Luminex bead-based methods, and patients subsequently developed antibody-mediated rejection of the donor kidney. All case reports were in female recipients with an HNA-3b3b genotype, who were sensitised to HNA-3a through pregnancy, transplantation, transfusion, or a combination thereof. Because the frequency of HNA-3b3b genotypes is low among the UK population at approximately 5%, some centres are pre-emptively screening females by HNA genotyping using PCR-SSP methods. Where an HNA-3b3b recipient is identified, anti-HNA antibody testing is performed.
Recently, a male HNA-3b3b patient with HNA-3a alloantibodies was reported by Oxford University Hospitals (Docker et al., 2024) with a failed allograft 11 years post-transplant. The patient’s genotype was determined through retrospective testing and the sensitising event was a blood transfusion. This may raise a question over screening strategies where only female patients are genotyped. Determining the patient and donor genotype could help identify whether anti-HNA antibodies detected are clinically relevant in avoiding adverse outcomes such as non-HLA antibody-mediated rejection.
Detection of HNA Alloantibodies
HNA alloantibodies can be detected with a range of in-house serological methodologies, such as agglutination, immunofluorescence, chemiluminescence, and bead-based Luminex assays. Granulocyte immunology results are usually inferred by looking for patterns of reactivity across a panel of donor granulocytes with known HNA types. Using a similar workflow to anti-HLA antibody detection, a bead-based assay by One Lambda can also be employed.
Granulocyte Agglutination Test (GAT)
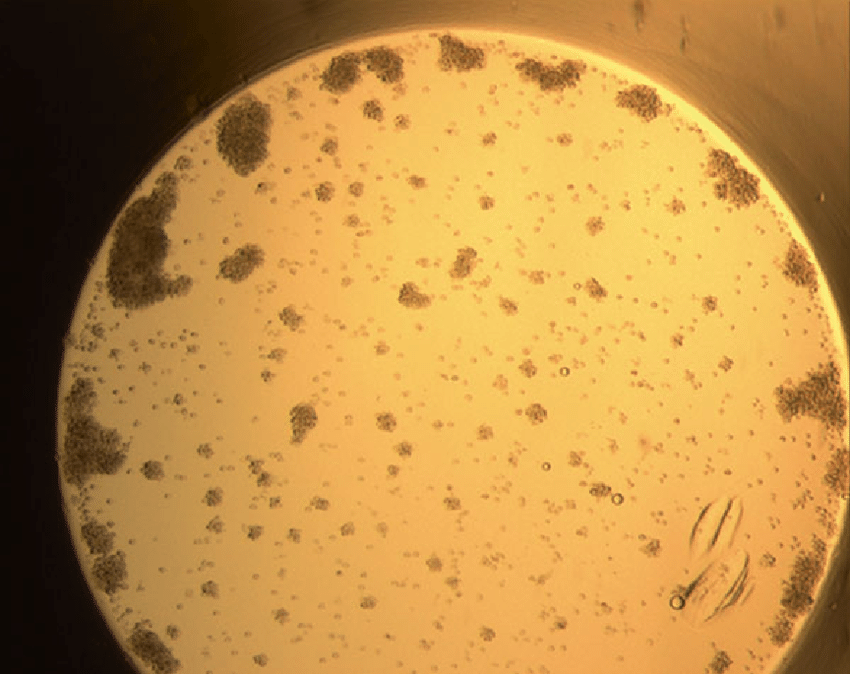
Fig. 1. Granulocyte agglutination test: Positive reaction with anti-HNA-3a typing serum (Dunn, 2015).
Granulocytes are incubated with patient serum. If antibodies are present, they will agglutinate.
Granulocyte/Lymphocyte Immunofluorescence Test (GIFT/LIFT)
More sensitive than GAT, a GIFT also incubates patient serum with fixed donor granulocytes and lymphocytes. Detection of alloantibodies is by flow cytometry using a fluorescently-labelled secondary IgG or IgM antibody. A direct GIFT can detect autoantibodies bound to the surface of the patient’s own granulocytes, while an indirect GIFT detects circulating autoantibodies in the serum of the patient that bind with donor granulocytes.
Granulocyte Chemiluminescent Test (GCLT)
A functional assay using unfixed, heat-treated granulocytes and lymphocytes incubated with patient serum. Monocytes are added, resulting in the phagocytosis of bound antibodies. This results in an oxidative burst and fluoresces in the presence of Luminol.
Monoclonal Antibody Immobilisation of Granulocyte Antibody (MAIGA)
MAIGA can be used to detect allo- or auto-antibodies against known HNA epitopes. Monoclonal antibodies are coated on a plate and used to immobilise, or capture, HNA expressed on fresh or frozen granulocytes. The patient’s serum is then added to the plate and HNA alloantibodies in the serum will bind the captured granulocyte antigens. The cells are solubilised, a substrate added, and a colorimetric change (from clear to yellow) is read using a tray reader, indicating the presence of antibodies (e.g. CD16 monoclonal antibodies may indicate HNA-1b1b specific antibodies). This is currently the gold standard method for identifying granulocyte antibody specificity.
LABScreen™ Multi
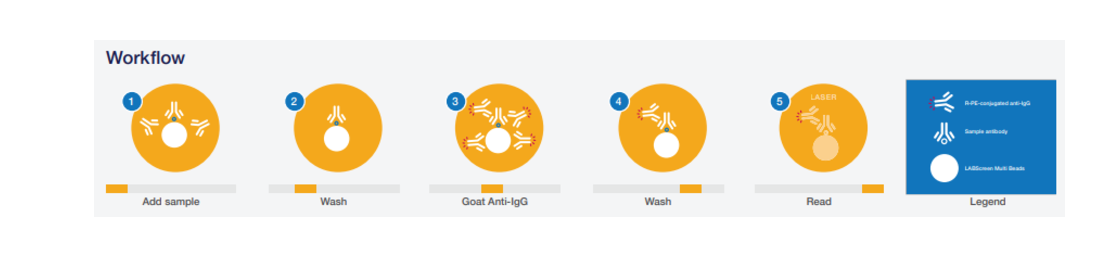
A multiplexed, kit-based test, employing a familiar workflow using Luminex xMAP® technology to simultaneously detect antibodies against HLA Class I, HLA Class II and HNA-1a, -1b, -1c, -2, -3a, -3b,- 4a, -5a, and -5b epitopes in a single well. This is a highly sensitive CE-IVD certified product. Microspheres or beads, uniquely identifiable by their internal dyes, are coated with known HLA and HNA antigens. Serum is incubated with the beads, before being washed by either centrifuging the plate or using a filter plate and vacuum manifold. The antibody-antigen complexes on the beads are incubated with a secondary fluorochrome-conjugated anti-human IgG antibody before a further wash step to remove unbound antibody. Beads are resuspended in phosphate buffered saline before being acquired on a Luminex 100/200 or FLEXMAP 3D instrument.
During acquisition, sheath fluid forces beads into single file as they pass through a cuvette, green and red lasers excite the fluorochrome attached to the antigen-antibody complex. Fluorescence and light scatter are detected by optics and used to identify the beads and quantify a median fluorescence intensity value for each bead. Whether the MFI value is above or below a threshold for each analyte will determine whether it is positive or negative.
HNA Genotyping
Table 1 outlines the five HNA systems, as defined by the International Society of Blood Transfusion (ISBT) HNA nomenclature subcommittee and the ISBT Granulocyte Immunology Working Party (GIWP). Single nucleotide polymorphisms in known positions can result in a different genotype and, therefore, phenotype. HNA genes FCGR3B, SLC44A2, ITGAM, and ITGAL are inherited in Mendelian fashion from each parent and are known to encode epitopes HNA-1a/b/c/d/bv/null; HNA-2/null; HNA-3a/b/3av; HNA-4a/b; and HNA-5a/bw. These epitopes are expressed on the following glycoproteins: Fc gamma receptor IIIb, CLT2, CD11b.CD18 and CD11a.CD18, respectively.
HNA genotypes can be multiallelic (e.g. a patient may have an FCGR3B*01,*02,*03 genotype and HNA-1a1b1d phenotype), or biallelic (e.g. an ITGAM*01*02 genotype encoding an HNA-4a4b phenotype), as is the case for HNA-3, -4, and -5. HNA-2 is not expressed on glycoproteins therefore is not included in routine HNA genotyping.
HNA genotyping can be performed using commercial methods, such as polymerase chain reaction using sequence-specific primers (PCR-SSP), PCR using sequencing-based technology (PCR-SBT), or real-time PCR (qPCR).
PCR-SSP

The One Lambda HNA Genotyping Trays target known polymorphisms for HNA-1, -3, -4 and -5 using primers that anneal complementary sequences flanking target regions. Sequence specific primers are lyophilised in wells, genomic DNA is added to wells with PCR reagents, and the mixture is transferred to a thermal cycler for the PCR reaction to take place. Amplicons are run on an agarose gel by gel electrophoresis and visualised with an ultraviolet illuminator. Presence of bands at known molecular weights can be interpreted using a worksheet. This is a relatively quick, inexpensive and simple way to ascertain HNA genotypes, with results in less than three hours.
PCR-SBT

PCR-SBT employs similar sequence-specific primers, targeting known SNPs in both forward and reverse direction, before a second PCR cycle sequencing reaction targets amplicons in one direction only with the inclusion of ddNTPs (e.g. BigDye Terminator). Purification steps using paramagnetic beads capture DNA fragments of a desired length in size and gel column plates are used to to remove remaining PCR reagents and dye terminators. The purified products are then acquired on genetic analysers using capillary gel electrophoresis. As each incorporated dye terminator passes a laser, the fluorescence is associated with a known base. The fluorescence is converted to a base call, and a sequence of base calls can then be aligned with a known reference sequence using genetic software. Novel SNPs can be identified if they exist within the region being targeted by primers, but the clinical significance of these is not always known as these are rarely reported. Compared to PCR-SSP, Sanger-based methods are expensive, laborious and require highly technical staff to perform the assay, interpret the results, and maintain the stock of in-house PCR-SBT reagents.
Real-time PCR
Another commercially available option: employing sequence-specific primers to target known SNPs and Taqman hydrolysis probes for a quantitative PCR-based approach. Fluorescence is detected by a real-time PCR machine and the results are analysed using software.
VH Bio Services and Support
As One Lambda’s licensed channel partner in the U.K. and Republic of Ireland, V.H. Bio Limited supply CE-IVD certified products and equipment for anti-HNA antibody identification (LABScreen™ Multi; 03LSMUTR) and HNA genotyping by PCR-SSP (HNA Genotyping Tray; 03HNAGEN).
References
Enquire about this article
"*" indicates required fields
