Image Credit: National Science Foundation, via Wikimedia Commons
Current challenges in transplantation
In organ donation and transplantation, our primary concern is to treat the patient whose organ is failing and, if doing so with a transplant, avoid acute rejection. For many patients, transplantation is the single best – or only – treatment option; yet, for more than 1 in 10 patients transplanted in the UK, their allograft fails within the first five years, and adverse events related to under- or over-immunosuppression continue to pose a significant challenge (1). There has been a long history of policies to increase donation and transplantation rates, e.g. the introduction of “deemed consent” in 2020 and the Organ Donation and Transplantation 2030: Meeting the Need strategy; however, there is no clear national policy for ensuring these donor organs last the test of time.
2025 has been a pivotal year in transplantation, with advances in molecular diagnostics like cell-free DNA and transcriptomics, AI, machine learning, and sustainability featuring heavily at BTS and ESOT congresses. At both events, NHSBT highlighted the persistent gap between transplant demand and donor organ availability. Notably, research presented at ESOT explored alternative organ sources, such as xenotransplantation and 3D-printed organs, while Sir Stephen Powis – then NHS England National Director – presented Fit for the Future: 10 Year Health Plan for England, aiming to reduce transplant demand by tackling root causes of organ failure, such as obesity, cardiovascular disease, and diabetes through health promotion. He also shared important insights into the sustainability of the NHS workforce (2).
Kidney Research UK’s (KRUK) comprehensive report on the health economics of kidney disease to 2033 highlighted some concerning findings:
- 1 in 10 people in the UK suffers from chronic kidney disease.
- 30,000 patients with renal failure in the UK are on dialysis.
- At a cost of £34,000 per patient per year, the annual burden of dialysis on the NHS is £7billion.
- The number of patients on dialysis could more than quadruple to 143,000 by 2033; 12,000 of whom are estimated to need a kidney transplant (3).
This trajectory is unsustainable – consideration must be given to prolonging allograft survival and preventing kidney transplant patients from returning to dialysis. This will require a multipronged, evidenced-based, consensus-driven approach; one that integrates cutting-edge technology, personalised care, and digital innovation (Figure 1).
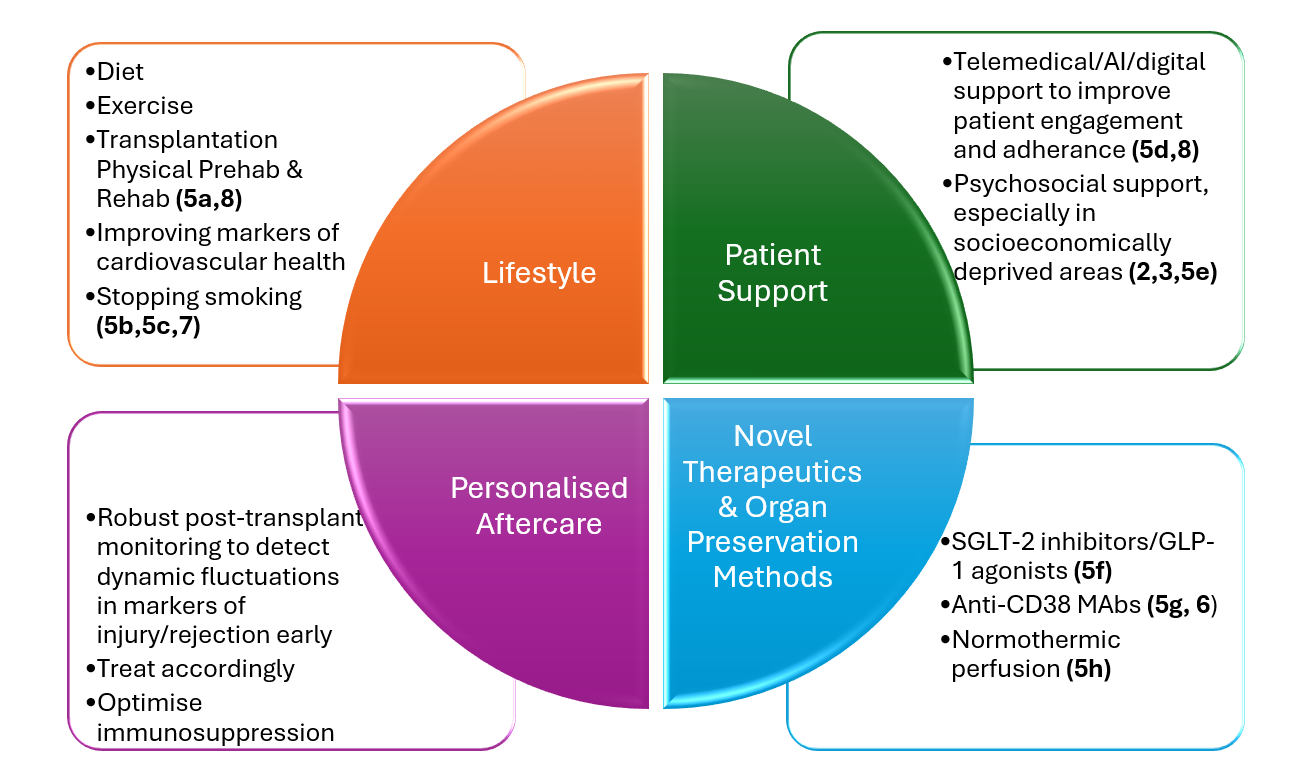
Figure 1. A holistic concept for improving transplant outcomes, including examples presented at ESOT Congress. MAbs= Monoclonal antibody targeted therapy; SGLT2= sodium-glucose cotransporter2; GLP-1=glucagon-like peptide-1 (5)
Current challenges in post-transplant monitoring
Following organ transplantation, methods of allograft surveillance in the UK vary depending on the organ(s) transplanted. After a heart transplant, allograft surveillance could look like 2-weekly biopsies, and multiple biopsies thereafter for up to two years. This is a daunting commitment to long hospital stays and regular invasive investigations, which can negatively impact patient wellbeing and risk damaging the graft itself. Although biopsy remains the gold standard for diagnosing graft injury and rejection after transplantation, its invasive nature makes it unsuitable for ongoing monitoring. Biopsy is also vulnerable to sampling errors and can yield ambiguous or conflicting histopathological results.
Following a kidney transplant, post-transplant monitoring usually involves regular blood and urine tests to establish kidney function, as well as monitoring whether levels of immunosuppressants are within therapeutic levels or drifting into cytotoxic levels. If rejection is suspected, biopsies and retrospective testing for the development of donor-specific antibodies are performed. Current biochemical markers of kidney function (e.g. eGFR and proteinuria) can lack specificity and be a late indicator of rejection, at which point interventions begin to lose their efficacy (9).
They can also be inaccurate. Take Tacrolimus monitoring for example: initial dosing is body weight dependent; what may be optimal for one person may not be for another; individual genetics and metabolism can affect clearance and subsequent levels in the blood, even in the same person over time. With such a tight therapeutic window, over-immunosuppression leaves the patient at risk of adverse effects of opportunistic infections and malignancies, while under-immunosuppression increases the risk of rejection (10).
In 1991, the Banff Classification was developed to standardise renal allograft pathology diagnoses and is now considered the international gold-standard approach. Two-yearly iterations of the Banff Classification are released following consensus, evidenced-based, expert review. Banff Classification historically lacked the nuance to diagnose borderline or suspected rejection cases; however, additional categories are added with each review alongside developments in diagnostic testing and, since 2017, there has been more of a focus on ever-evolving molecular diagnostic methods (Figure 2; 11).
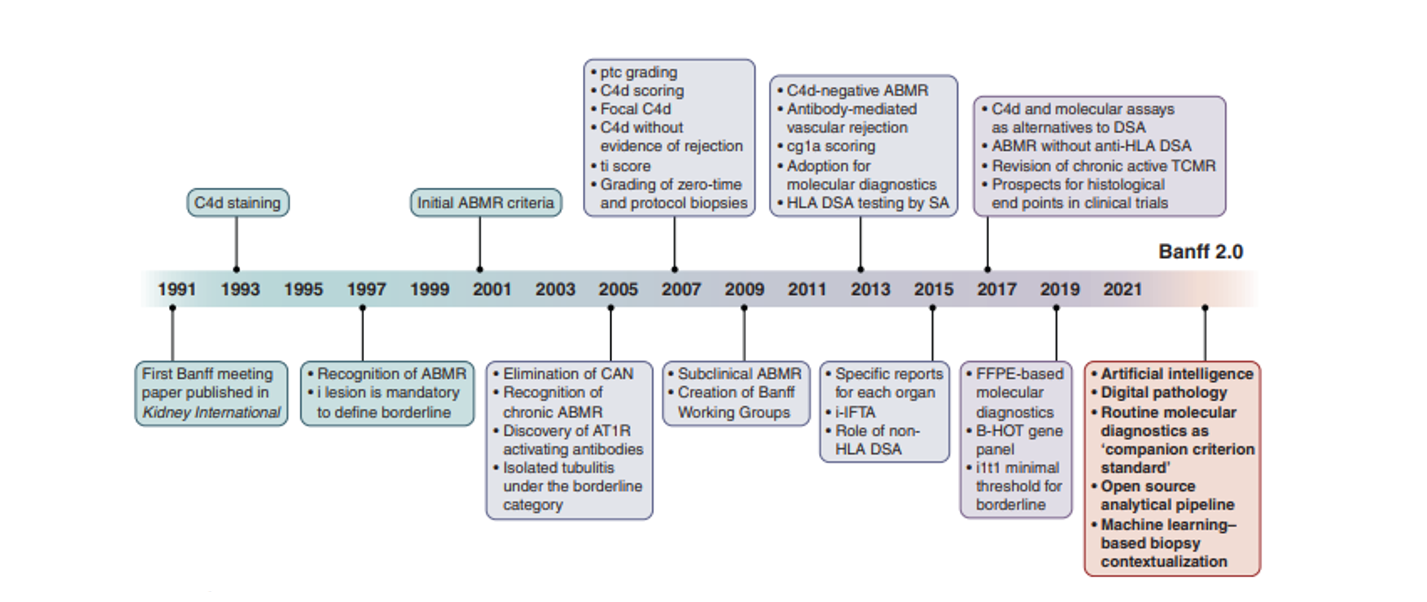
Figure 2. 30-Year Evolution of the Banff Classification: notable changes and future concepts. ABMR=antibody-mediated rejection; AT1R=angiotensin type 1 receptor; B-HOT=Banff-Human Organ Transplant; CAN=chronic allograft nephropathy; cg= transplant glomerulopathy; DSA=donor-specific antibody; FFPE= formalin-fixed paraffin-embedded; HLA= human leukocyte antigen; i=interstitial inflammation; i-IFTA= inflammation within areas of interstitial fibrosis and tubular atrophy; ptc=peritubular capillaritis; SA= single antigen; t= tubulitis; TCMR=T cell–mediated rejection; ti= total cortical inflammation (Adapted from Loupy et al; 11)
Emerging markers with potential to transform patient management
Donor-Specific Antibodies (DSAs)
In 2011, a prospective study by Sellares and colleagues looked to further define rejection subtypes using biopsy diagnoses, HLA antibodies and clinical data. The leading cause of rejection in their cohort was ABMR with nearly half of all rejection losses observed in non-adherent patients, highlighting the importance of both antibody and immunosuppression monitoring (12). NHS Blood and Transplant (NHSBT) data from over 22,000 transplanted kidneys between 2000-2013 revealed that a staggering 1 in 5 allografts failed within the first 5 years, with alloimmune pathology being the leading cause (25%) of allograft failure, after death with a functioning kidney (13). Although allograft failure rates have improved drastically to just 1 in 10 at 5 years, according to recent NHSBT activity reports, graft loss at this early stage is undesirable (14).
Pre-transplantation, patients on the transplant waiting list are screened for pre-existing anti-HLA antibodies. HLA antigens deemed to be ‘unacceptable’ – those with antibody titres likely to cause a positive crossmatch and veto the transplant – are reported to the national organ allocation team and avoided. This process is crucial in preventing unnecessary shipment of donor organs to patients with an immunological risk of hyperacute rejection, which can result in early graft loss and be life-threatening.
Post-transplantation, DSAs can be produced by the recipient’s immune system in response to mismatched HLA on the allograft, in blood transfusions, or through pregnancies. The development of persistent de novo DSA (dnDSA) is strongly associated with antibody-mediated rejection (ABMR) and chronic rejection (OuTSMART trial; 15).
Despite being a sensitive marker of subclinical and clinical ABMR, post-transplant monitoring of DSAs is not routinely performed by many transplant laboratories in the UK and Republic of Ireland and is often only investigated for-cause.
Detection: dnDSA can be detected using bead-based assays, where microscopic, polystyrene beads are coated in HLA antigens to which circulating anti-HLA antibodies can bind (Figure 3). A secondary antibody (PE-conjugated anti-human IgG) binds to the antibody-antigen-bead complex, and the fluorescence is detected and quantified using a Luminex instrument. Software is used to analyse the pattern of reactivity against known bead IDs and local reactivity thresholds, and results are verified by a trained scientist. A patient’s antibody profile can be more fully characterised using single-antigen bead (SAB) and extended panel assays. LABScreen™ Single Antigen Bead assays are highly sensitive and specific for HLA antibodies (allele-level), which has led to the phasing out of more laborious, cell-based methods, such as the complement-dependent cytotoxicity crossmatch.
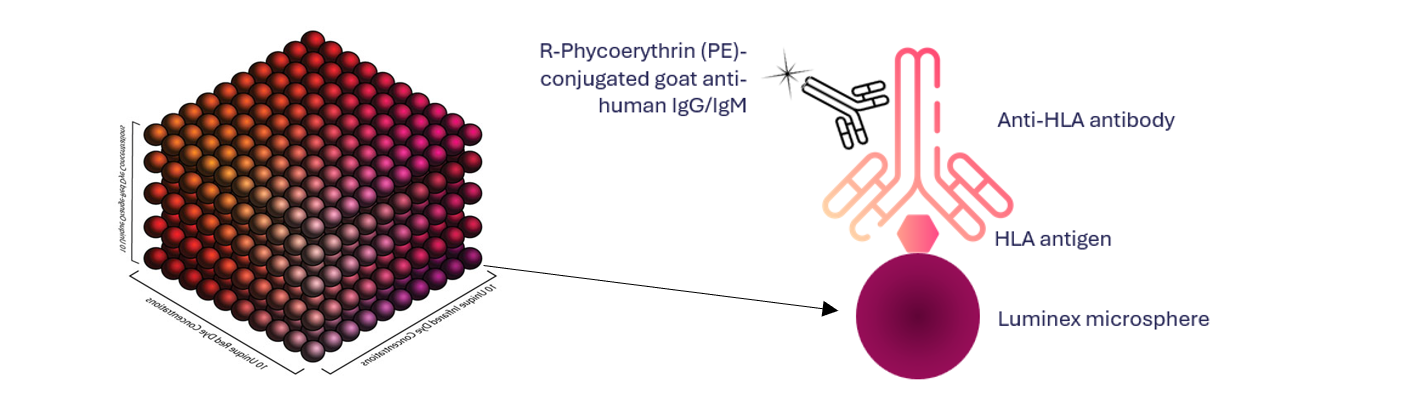
Figure 3. Luminex microbeads are coated with Human Leukocyte Antigens (HLA). Anti-HLA antibodies in patient serum bind HLA on the bead surface. A secondary fluorescent antibody binds to the antibody-antigen-bead complex. Red laser distinguishes Bead ID using internal dyes, while green laser detects fluorescence (MFI) of the secondary antibody.
Considerations: Not all DSA-positive patients develop rejection and biopsy-proven ABMR can occur in patients without DSA, e.g. due to anti-HNA antibodies. Bead-based assays are solid phase, so MFI values are semiquantitative and should be interpreted by experienced scientists in the context of the patient’s clinical history.
Clinical Utility: Regularly testing serum samples post-transplant could identify patients with persistently high MFI dnDSA who may be at greater risk of adverse graft outcomes; especially if directed against HLA-DQ (15, 16).
Who will benefit? All solid organ transplant recipients.
Donor-Derived Cell-Free DNA (dd-cfDNA)
Following apoptosis or necrosis of the donor organ cells, cell-free DNA enters the patient’s circulation. Dd-cfDNA can be isolated from plasma and quantified (as copies/mL or as a percentage of the total cfDNA). With a negative predictive value of 95%, unnecessary biopsies can be avoided in patients showing graft quiescence. Elevated levels of dd-cfDNA can indicate graft injury, often preceding clinical signs of rejection. A dd-cfDNA level ≥0.12% has been described as clinically relevant in heart transplantation (17), while a rise in dd-cfDNA of ≥0.5% strongly correlates with both clinical and subclinical rejection with elevations in dd-cfDNA, preceding DSA detection by a median of 91 days, and enabling a diagnosis of ABMR as early as 12 months ahead of standard care (18,19). Whether low or high, this demonstrates the importance of highly sensitive methods and longitudinal monitoring for these small increases from a patient’s baseline. ESOT’s 2024 Consensus Statement on Testing for Non-Invasive Diagnosis of Kidney Allograft Rejection states:
“…that clinicians consider measuring serial plasma dd-cfDNA in patients with stable graft function to exclude the presence of subclinical antibody mediated rejection.” (20)
Furthermore, dd-cfDNA has a short half-life; levels may be utilised as a dynamic marker to monitor graft health in response to therapeutic interventions or adjustments.
Detection: dd-cfDNA can be isolated from human plasma and quantified by sensitive molecular methods such as next generation sequencing (NGS; Fig. 4) or digital droplet PCR and monitored over time. Importantly, Dr Philip Halloran’s Trifecta group demonstrated that these methods yield comparable results (21), meaning clinicians and laboratories can choose a platform that suits their existing infrastructure and consider practical utility without sacrificing diagnostic accuracy.
The One Lambda Devyser Accept cfDNA assay identifies donor-specific “informative markers” pre-transplantation which can be monitored post-transplantation, improving specificity. Given approximately 1 in 5 patients ends up being re-transplanted, it is also useful to know that Accept cfDNA is validated for monitoring dd-cfDNA from more than one donor (22).
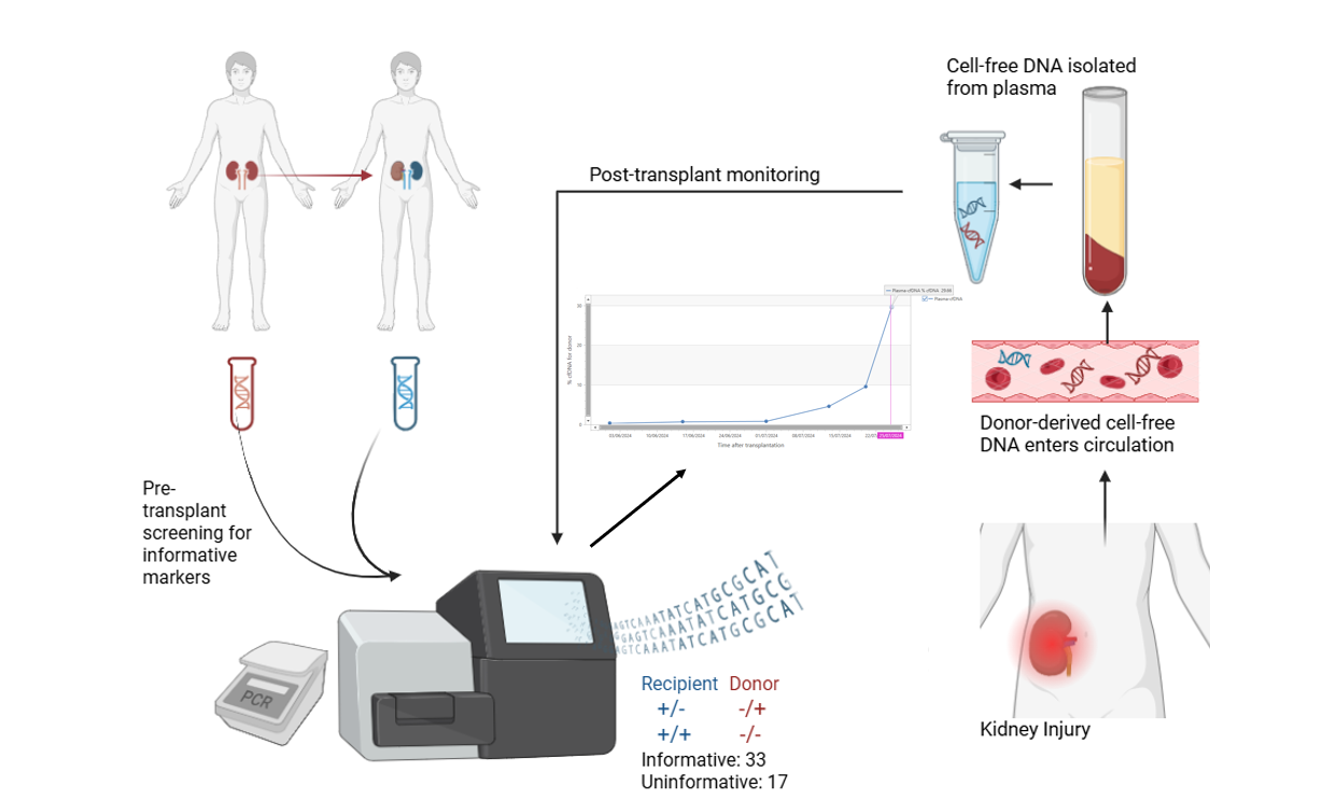
Figure 4. Donor-derived cell-free DNA monitoring using One Lambda Devyser Accept cfDNA.
Considerations: Though the limit of detection of dd-cfDNA is highly sensitive – as low as 0.1% for Accept cfDNA – dd-cfDNA is unable to distinguish cause of rejection in isolation. Given the short half-life of cfDNA, specialist preservation tubes are also recommended.
Clinical Utility: Elevated levels can be detected at least 3 months ahead of dnDSA and 6 months ahead of a rise in serum creatinine or decline in eGFR. Elevated dd-cfDNA predicts a three-fold increase in eGFR and enables earlier diagnosis of ABMR, providing an earlier window of opportunity for therapeutic intervention (6, 18, 19). Elevated dd-cfDNA strongly correlates with ABMR, T-cell mediated rejection (TCMR), and mixed rejection, according to a study by Aubert et al. of over 1,000 patients in Paris (23).
After reviewing the evidence, the American Society for Transplant Surgeons (ASTS) issued a position statement saying that dd-cfDNA shows significant utility in diagnosing rejection earlier than standard care, while dd-cfDNA <0.21% showed a NPV of 95%, negating the need for biopsy (24, 25).
Rising dd-cfDNA in an otherwise well patient may indicate allograft injury, e.g. calcineurin inhibitor toxicity, and signal that further investigation may be required. Recent UK pilot studies have confirmed these findings (26,27). Dd-cfDNA has proven useful in diagnosing microvascular inflammation in C4d negative and DSA negative patients (28). Combining dd-cfDNA with dnDSA and gene expression increases their combined positive predictive value, outperforming traditional histology in predictive power (29).
Who will benefit? One Lambda Devyser Accept cfDNA is currently CE-IVD/IVDR certified for kidney transplant recipients; studies are underway for other solid organs.
Urinary Chemokines
C-X-C motif chemokine ligand-10 (CXCL10) is a pro-inflammatory chemokine stimulated by interferon gamma; hence, it is also known as interferon gamma-induced protein 10 (IP-10). Urinary CXCL10 can be significantly elevated during acute rejection or subclinical graft injury, so has emerged as a promising marker of kidney allograft rejection as it is organ-specific and can be detected without the need for invasive procedures (30).
Detection: Urine is an ideal, non-invasive sample source. Currently performed by ELISA methods by some research groups, urinary CXCL10 is also available as a Luminex bead-based assay at a ThermoFisher Scientific service lab in the US. Using this test, if present in the urine, CXCL10 can be captured by IP-10-coated capture beads, and fluorescently ‘tagged’ with a secondary streptavidin R-phycoerythrin conjugate (SAPE) antibody, before analysing on a Luminex instrument. This may, in future, become a routine diagnostic assay for clinical laboratories (31).
Considerations: In isolation, elevated urinary CXCL10 is not diagnostic for rejection, and it is less sensitive than dd-cfDNA at detecting subclinical rejection (32).
Clinical Utility: Elevated urinary CXCL10 is a non-invasive early indicator of graft dysfunction. Urinary CXCL10 may prove useful as a screening tool as part of a wider algorithm for assessing allograft pathology.
Who will benefit? Kidney transplant recipients.
Omics
For centuries, the pursuit of understanding human biology in both health and disease has propelled medical and scientific advancements – from structural and anatomical observations to intercellular interactions; each advancement has helped develop diagnostic and therapeutic strategies. In the 1960s, X-ray crystallography helped determine the structures of tens of thousands of proteins and nucleic acids. Proteomics explored protein structure and function by gel electrophoresis and mass spectrometry. Genetics focuses on individual genes, while genomics studies the entire genome, and how genes are expressed and interact with each other. The integration of data from multiple molecular disciplines, multiomics, is providing unprecedented insight into immune regulation, graft tolerance, and rejection mechanisms, as well as targets for potential therapeutics (see Table 1).
Table 1. Multiomics integrates multiple biological processes, each with their own “-omics”.
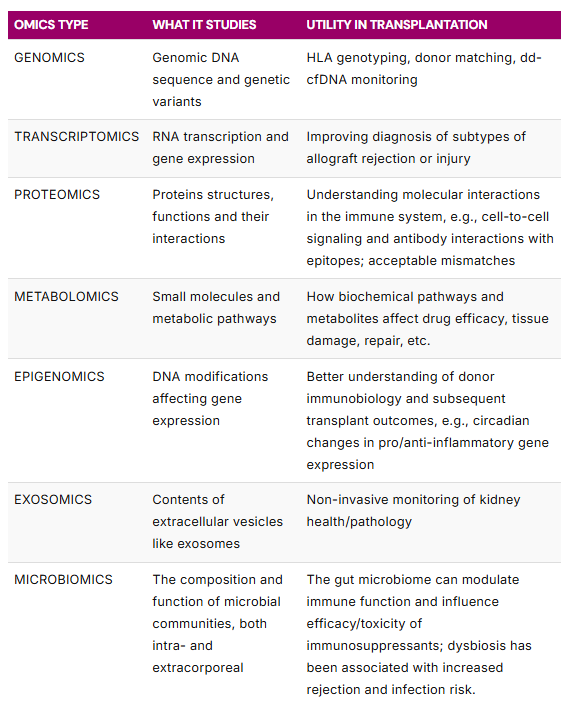
Transcriptomics is the study of the total set of RNA molecules, e.g. messenger RNA (mRNA) which drives protein synthesis, and microRNA (miRNA) which regulates gene expression. Pre-transplantation, analysis of RNA signatures in the blood, e.g. One Lambda’s Pre-Transplant Risk Assessment (PTRA) assay (developed by Basterd and colleagues and licensed by Verici Dx), can help risk stratify patients into high or low risk of early acute rejection in kidney transplant recipients, which could help inform patient management (33).
Post-transplantation, transcriptomics can be studied using single cell sequencing methods like Illumina and NanoString, while microarray can be used to analyse gene expression across hundreds to thousands of genes simultaneously. By analysing the total set of RNA and seeing which genes are expressed, a “gene expression profile” can be determined.
MicroRNAs (miRNAs) are small, non-coding RNAs that regulate gene expression by altering the translation of their target mRNAs. MiRNAs are critical regulators of homeostasis, so their dysregulation is an important component of cell and organ injury. A growing body of evidence shows that miRNAs are involved in the pathophysiology of acute kidney injury (AKI), chronic kidney disease (CKD) and allograft damage, with different subsets becoming dysregulated depending on the condition; therefore, they have utility in graft surveillance (34).
Prof. Menna Clatworthy’s Group at the University of Cambridge, has been analysing gene expression using single cell RNA sequencing data from foetal and adult kidneys to generate a world-first “human kidney cell atlas” as part of the Human Cell Atlas project. This work has revealed molecular signals by epithelial cells in regions which need protecting from infection and could help identify cellular targets for drug therapeutics (35).
Jaffree and colleagues published works recently, combining single cell RNA sequencing with powerful 3D imaging to “map” lymphatic vessels in donor kidney tissue, demonstrating changes in lymphatic vessel ‘mapping’ during chronic rejection (Figure 6; 36).
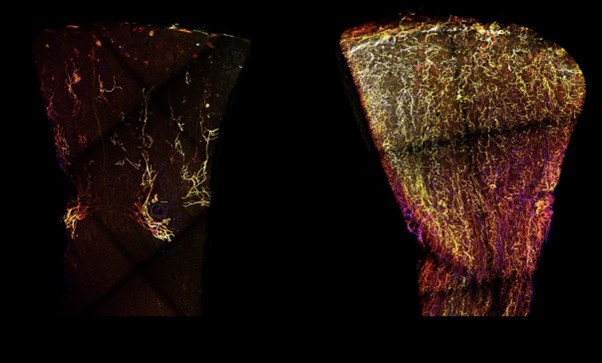
Figure 5. The lymphatic vessels of a healthy kidney (left) are represented in yellow, compared to kidney undergoing rejection (right). Jaffrey et al (2025) [Image from https://tinyurl.com/aenuws5n accessed 03/10/25]
After performing a literature search of peer-reviewed microarray and NanoString publications on transplanted organs, the Banff Molecular Diagnostics Working Group compiled a list of validated transplant outcome-related genes of interest which formed the Banff-Human Organ Transplant (B-HOT) set of 758 genes of interest. In the UK, Beadle and colleagues compared transcriptomics of kidney biopsies with suspected ABMR and found that their 9-gene score was highly predictive of active ABMR and strongly associated with risk of allograft loss (37).
The Molecular Microscope Diagnostic System (MMDx)
Since 2006, Dr. Philip Halloran’s group in Canada has been pioneering transcriptomics using microarray technology to develop the concept of pathology-associated transcript sets in allografts (INTERCOMEX study). MMDx uses machine learning to compare the expression of thousands of RNA transcripts in biopsy tissue to a reference set of hundreds of previously characterised biopsies from transplanted patients to identify patterns associated with TCMR, ABMR, and parenchymal injury (e.g. acute kidney injury or fibrosis). Unlike traditional histology, which relies on visual interpretation and can be subjective, MMDx offers objective, reproducible analyses of graft health and shows better agreement with clinical opinion compared to traditional histology. It can be used to detect subclinical rejection and injury not visible by microscope, to complement or clarify ambiguous histology findings, and to provide further information to clinicians regarding allograft injury severity (38).
Detection: Samples are stored in RNALater and shipped at ambient temperature. Total RNA is extracted from biopsy samples, labelled, hybridised to a microarray chip, washed and stained, then the chips are read. Gene expression of 49,000 transcripts is interpreted using a predefined machine learning algorithm to distinguish graft injury subtypes.
Considerations: Sites licenced to perform MMDx are currently limited to: Canada, USA, and Czech Republic. MMDx still requires (small amounts of) biopsy tissue and can be costly. In 2024, there was insufficient evidence for ESOT or ASTS to make consensus recommendations for the use of gene expression profiling; however, this is a fast-evolving area of research with many clinical trials underway (Trifecta-Lung, Trifecta-Heart, Trifecta-Kidney, among others).
Clinical Utility: MMDx represents a shift toward precision diagnostics in transplantation, offering a deeper understanding of graft status, helping tailor immunosuppressive strategies more effectively, while providing enhanced agreement with clinicians compared to histological findings. MMDx has shown strong correlations with DSA and dd-cfDNA levels, enhancing its diagnostic power of ABMR, TCMR and mixed rejection when used alongside these markers.
Who will benefit? MMDx is available as a lab service through ThermoFisher Scientific and their partner labs for kidney, heart and lung transplant recipients.
Urinary Exosomes
Exosomes are tiny, extracellular vesicles that play an important role in cell-to-cell interactions; how they are formed and their contents encapsulated is depicted in Figure 5. Studies have shown that exosomes from the kidney are secreted into the urine, and their molecular and biological contents can be reflective of kidney health or pathology. Transcriptomics has revealed miRNA transcripts, or signatures, that could differentiate patients with different nephropathies, such as IgA nephropathy, membranous nephropathy, lupus nephritis, paraprotein-related nephropathies, and diabetic nephropathy, as well as autosomal dominant polycystic kidney disease, from healthy controls (39).
Post-transplantation, urinary exosomal mRNA profiles can help differentiate ABMR from TCMR with high positive and negative predictive values (93.3% and 86.2%, respectively).
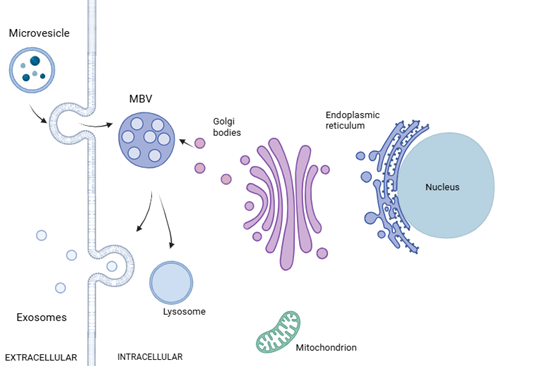
Figure 6. The formation of urinary exosomes begins with the cellular plasma membrane budding inwards, maturing into endosomes, which bud into vesicles and multivesicular bodies (MVBs), encapsulating proteins, DNA, messenger RNA, microRNA, and other biologically active molecules in the cytoplasm. Following fusion with the cell membrane, MBVs can be released into the extracellular space as exosomes, or fuse with lysosomes in the cell and undergo degradation MBV=multivesicular bodies
Detection: Urinary exosomes can be collected, transported and stored at 4°C. Thanks to their bilipid membranes, they are stable when stored chilled. Following isolation, urinary exosomal mRNAs can be evaluated by qPCR (currently a One Lambda laboratory service test).
Considerations: While sensitivity is 94%, the positive predictive value is 62%.
Clinical Utility: Urinary exosomes can offer insights into allograft function. Presence of certain mRNA signatures may aid diagnosis of rejection or inflammation, distinguish TCMR from ABMR, and potentially serve as a prognostic marker (40).
Who Will Benefit? Kidney transplant recipients
Torque Teno Virus (TTV)
Taking a slight detour from markers of graft injury, Torque teno virus is a non-pathogenic, non-enveloped, highly polymorphic human virus belonging to the Alphatorquevirus genus of the Anelloviridae family of viruses. TTV is thought to be extremely prevalent, uniformly represented in 65-100% of the human population across the globe. TTV does not have an etiological role like other viruses and its replication is controlled by the immune system, making it a potential surrogate marker of functional immune status, which can help guide personalised immunosuppression (41).
Detection: DNA for TTV can be extracted from whole blood or plasma. One Lambda (ThermoFisher) have developed an assay to quantify TTV by qPCR and digital PCR, with reproducible and linear results observed between the two platforms (41).
Clinical Utility: High TTV load may indicate over-immunosuppression, increasing risk of infection. Low TTV load suggests under-immunosuppression, potentially leading to rejection. High TTV and overimmunosuppression has also been associated with an increased risk of malignancies after kidney transplantation (Fig. 7; 42; 5i).

Figure 7. How Torque teno virus (TTV) load informs immunosuppression levels
Considerations: TTV levels may vary across different age groups and immunosenescence in older age may be a confounding factor. More high-quality research is needed to fully elucidate its clinical utility; the TTVguideTX trial looks set to do so (43).
Who will benefit? All solid organ transplant recipients on immunosuppression.
Translation of technological innovations into clinical practice
Integrating these molecular markers in the form of multiomics shows enormous potential for post-transplant patient management and care.
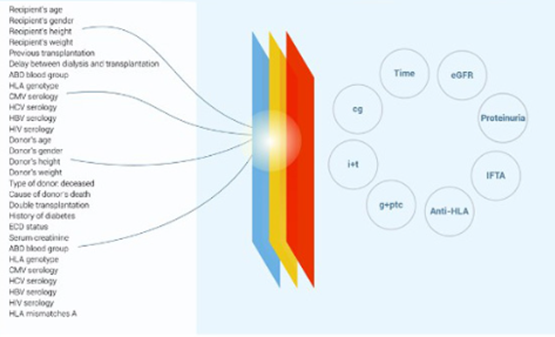
Figure 8. The Paris Transplant Group’s iBox algorithm
Using AI and machine learning to analyse results and identify patterns, algorithms like “iBox” (Figure 9) can not only consider pre-transplant variables, such as donor quality, virology results, routine biochemical markers, and pre-existing anti-HLA antibodies, but also post-transplant markers to continuously stratify risk of rejection, engage patients in their own care and transplant trajectory, and evaluate treatment options, to deliver truly personalised care.
Exciting news for transplant recipients in France, so where does that leave patients in the UK & RoI?
If, as with ESOT and the American Society for Transplant Surgeons, national consensus emerges regarding the clinical utility of these molecular markers, a unified, national strategy will be essential. This would include validation of post-transplant monitoring assays, machine learning to find patterns associated with graft injury or rejection, and consideration of an integrated, actionable report for clinicians. Prospective clinical trials will be needed to evaluate how these markers guide further testing (e.g. dd-cfDNA-directed biopsies) and therapeutic interventions (e.g. immunosuppression adjustments). Over time, an algorithm to guide testing will emerge (Figure 9). Health economics analyses will also be vital, with cost benefits reported elsewhere (44,45).
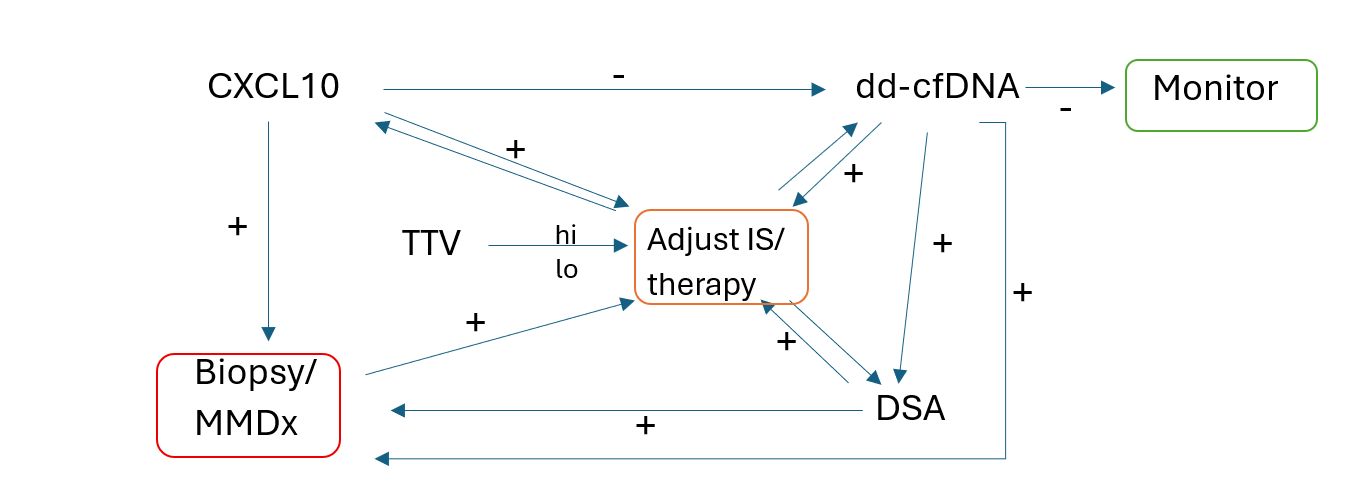
Figure 9. A post-transplant monitoring algorithm concept.
The pathway to implementation of technological innovations hasn’t always been clear; however, through national hubs like the NIHR HealthTech Research Centre in Leeds, whose research areas include “early diagnosis and personalised care” and “minimally invasive therapies and digital technologies”, charitable organisations like KRUK investing in research (e.g. Nita et al, 2025), and the UK government’s Life Sciences Sector Plan hoping to accelerate health innovations to deliver better outcomes for patients, strides forward can be made. Gauthier and colleagues from the Trifecta study group beautifully presented the relationship between dd-cfDNA, DSA, and gene expression (21). Using DART, ADMIRAL, Trifecta and the Paris Transplant Group’s studies as blueprints, the UK could completely transform NHS post-transplantation management and care and, hopefully, change the trajectory of patients needing dialysis for future generations.
Final Thoughts
These are transformative times in transplantation. With the convergence of molecular diagnostics, AI, and personalised medicine, we have a real opportunity to redefine patient care, extend graft survival, and ease the economic burden on the NHS. The upcoming joint BTS BSHI Monothematic Meeting on dd-cfDNA and omics marks a critical milestone – one that could propel the UK’s journey from innovation to implementation.
The tools are here, the evidence is growing… The rest is up to you.
If you would like to find out more about any of the information covered in this article, please get in touch with the Transplant Team at VH Bio.
Abbreviations
ABMR – Antibody-Mediated Rejection
ADMIRAL – Assessing Dd-cfDNA Monitoring Insights of Renal Allografts with Longitudinal Surveillance
BSHI – British Society for Histocompatibility and Immunogenetics
BTS – British Transplantation Society
CXCL10 – C-X-C Chemokine Ligand-10
DART – Circulating Dd-cfDNA in blood for Diagnosing Acute Rejection in kidney Transplant recipients
dd-cfDNA – Donor-derived cell-free DNA
DSA – Donor Specific Antibody
ESOT – European Society for Organ Transplantation
HLA – Human Leukocyte Antigen
IS – Immunosuppression
KRUK – Kidney Research UK
MFI – Median Fluorescence Intensity
MMDx – Molecular Microscope Diagnostic System
NGS – Next Generation Sequencing
NHSBT – NHS Blood and Transplant
NICE – National Institute for Health and Care Excellence
NIHR – National Institute for Health and Care Research
PTG – Paris Transplant Group
SAB – Single Antigen Beads
TCMR – T-cell-Mediated Rejection
TTV – Torque teno virus
References
1. NHSBT Activity Report 2024/25: Kidney
2. Fit for the Future: 10 Year Health Plan for England
4. Kidney Care UK Policy Updates
5. Abstract Book 22nd Congress of the European Society for Organ Transplantation
a. Physical Prehabilitation and Rehabilitation: BOS_11_15 Lloyd et al. Integrating Enhanced Recovery After Surgery in Renal Transplant Patients: Physiotherapy Perspective; PP-228 Leunis et al. Pre-Transplant Sarcopenia and its Progression Post-Liver Transplant: Impact on Long Term Survival
b. Smoking: BOS60_04_4 Ahmed, H. et al. Health Related Quality of Life in the First Year Post Kidney Transplantation – A Cohort Study; BOS_10_4 Beaudre, T. et al. Incidence, Risk Factors and Impact of Coronary Artery Disease After Kidney Transplantation; EP_170 Koch, M. et al. Surgical Complications of Living Kidney Donation with Different Surgical Techniques in Germany; EP_385 Maffei, I. et al. Can the Donor’s and the Recipient’s Lifestyle Influence the Development of Early CAV in the Recipient?
c. Telemedical: FOS_04_1 Mariat, C. et al. Environmental Impact of TeleMonitoring in Kidney Transplantation: An Ancillary Study of the Ap’Tx Clinical Trial (NCT03750331)
d. Digital Support EP_025: Walklin et al. Addressing Digital Exclusion from Physical Activity Digital Health Interventions: The EX-TAB Study (Kidney BEAM)
e. Psychosocial BOS60_04_4 Haupenthal, F. et al. Health Related Quality of Life in the First Year Post Kidney Transplantation – A Cohort Study
f. SGLT2/GLP-1: FOS_03_2 Rodriguez Espinosa, D. et al. SGLT2I Reduce the eGFR Slope in Diabetic and Non-Diabetic Renal Transplant Recipients; BOS_10_10: Cohen et al. Glucagon Like Peptide1 Receptor Agonists and Renal Outcomes in Kidney Transplants with Diabetes
g. Anti-CD38 Mab: FOS_10_1: Schatzl, M. et al. Monitoring Anti-CD38 Treatment with Felzartamab in Antibody-Mediated Kidney Allograft Rejection
h. Organ Perfusion: EP_386: Nieto, T. & Fedder, A. Normothermic Regional Perfusion Outcomes in Kidney Transplants at UHB
i. TTV: FOS_06_9: Herz, C.T. et al. Torque Teno Virus is Associated with an Increased Risk for Malignancies after Kidney Transplantation
6. Mayer, K. et al. (2024) A Randomized Phase 2 Trial of Felzartamab in Antibody-Mediated Rejection, N Engl J Med 391: 122-132
7. NICE Guideline (NG209) Tobacco: preventing update, promoting quitting and treating dependence [Accessed 16/10/25]
8. Greenwood, S.A. (2024) Evaluating the effect of a digital health intervention to enhance physical activity in people with chronic kidney disease (Kidney BEAM): a multicentre, randomised controlled trial in the UK, The Lancet Digital Health, 6 (1), e23-e32
9. Herath, S. et al.(2019) Advances in Detection of Kidney Transplant Injury. Mol Diagn Ther 23: 333–351
10. Francke, M.I. et al. (2020) The pharmacogenetics of tacrolimus and its implications for personalized therapy in kidney transplant recipients, Expert Review of Precision Medicine and Drug Development, 5:5, 313-316
11. Loupy, A. et al (2022) Thirty years of the International Banff Classification for Allograft Pathology: the past, present, and future of kidney transplant diagnostics, Kidney International, 101 678-691
12. Sellarés J, et al. (2012) Understanding the causes of kidney transplant failure: the dominant role of antibody-mediated rejection and nonadherence. Am J Transplant. 12 (2): 388-399
13. Burton, H. et al. (2019) Causes of renal allograft failure in the UK: trends in UK Renal Registry and National Health Service Blood and Transplant data from 2000 to 2013, Nephrol Dial Transplant 34: 364–370
14. NHS Blood and Transplant Annual Report on Living Donor Kidney Transplantation 2024/25
15. Jaffer, A. et al. (2025) Transplant outcomes after de novo donor specific antibody (DSA) development correlate with persistence of DSA, Transplant Immunology, 93
16. Tambur, A.R. et al. (2021) Significance of HLA-DQ in kidney transplantation: time to reevaluate human leukocyte antigen-matching priorities to improve transplant outcomes? An expert review and recommendations, Kidney Intl, 5 1012-1022
17. Moeller, C.M. et al. (2025) Clinical Utility of Combined Donor-Derived Cell-Free DNA and Peripheral Gene-Expression-Profiling in Heart Transplant Recipients, Clinical Transplantation, (39): 10 https://doi.org/10.1111/ctr.70340 [Accessed 16/10/25]
18. Bu, L. et al. (2022) Clinical outcomes from the Assessing Donor-derived cell-free DNA Monitoring Insights of kidney Allografts with Longitudinal surveillance (ADMIRAL) study, Kidney International 101:793-803
19. Akifova, A. et al. (2025) Donor-derived cell-free DNA monitoring for early diagnosis of antibody-mediated rejection after kidney transplantation: a randomized trial, Nephrol Dial Transplant, 40, 1384-1395
20. Park, S. et al. (2024) European Society of Organ Transplantation Consensus Statement on Testing for Non-Invasive Diagnosis of Kidney Allograft Rejection, Transpl Int 36:12115.
21. Gauthier, P.T. et al. (2024) Distinct Molecular Processes Mediate Donor-derived cell-free DNA Release from Kidney Transplants in Different Disease States, Transplantation 108 (4): 898-910
22. Pettersson, L. et al. (2024) Detection of donor-derived cell-free DNA in the setting of multiple kidney transplantations, Frontiers in Immunology, 15: 1282521
23. Aubert, O. et al. (2024) Cell-free DNA for the detection of kidney allograft rejection, Nat Med, 8 (8): 2320-2327
24. ASTS Position Statement 2024 Update
25. Bloom, R. et al. (2017) Cell-Free DNA and Active Rejection in Kidney Allografts, J A Soc Nephrol. 28 (7) 2221-2232
26. Nita, G. et al (2025) A novel INDEL-based next-generation sequencing assay for monitoring donor-derived cell-free DNA in renal transplant recipients – from bedside to results: a UK pilot study, Clin Transplant Res https://doi.org/10.4285/ctr.25.0004 [Accessed 16/10/25]
27. Clark, B. et al (2025) DdcfDNA Monitoring – A Single-Centre Pilot Study, Abstract, Int J Immunogenet, 52: S14
28. Aubert, O. et al. (2024) Cell-free DNA for the detection of kidney allograft rejection, Nature Medicine, 30 2320-2327
29. Halloran, P.F. et al. (2022) The Trifecta Study: Comparing Plasma Levels of Donor-derived Cell-Free DNA with the Molecular Phenotype of Kidney Transplant Biopsies, J Am Soc Nephrol, 33 (2) 387-400
30. Janfeshan, S. et al. (2024) Urinary CXCL-10, a prognostic biomarker for kidney graft injuries: a systematic review and meta-analysis, BMC Nephrol. 25 (1):292
31. Ho, J. et al. (2023) Multicenter Validation of a Urine CXCL10 Assay for Noninvasive Monitoring of Renal Transplants, Transplantation, 107 (7) 1630-1641
32. Abstract Book 22nd Congress of the European Society for Organ Transplantation BOS_14_17 Rodriguez, I.T. et al. Concominant Assessment of Urinary Chemokines and Donor-Derived Cell-Free DNA for the Diagnosis of Kidney Rejection
33. Basterd, O. et al. (2024) Prospective observational study to validate a next-generation sequencing blood RNA signature to predict early kidney transplantation, Am J Transplantation, 24, 436-447
34. Mahtal, N. et al. (2022) MicroRNAs in kidney injury and disease, Nat Rev Nephrol, 18 (10): 643-662
35. Stewart, B.J. et al. (2019) Spatio-temporal immune zonation of the human kidney, Science, 365 (6460) 1461-1466
36. Jafree et al. (2025) Organ-specific features of human kidney lymphatics are disrupted in chronic transplant rejection, J Clin Invest.; 135(18)
37. Beadle, J. et al. (2023) Application of the Banff Human Organ Transplant Panel to kidney transplant biopsies with features suspicious for antibody-mediated rejection, Kidney International 104 (3) 526 – 541
38. Halloran, P.F. et al. (2017) Real Time Central Assessment of Kidney Transplant Indication Biopsies by Microarrays: The INTERCOMEX Study, Am J Transplantation, 17: 2851-2862
39. Gaudio, C. et al. (2025) Urinary Exosomes in Nephrology: A New Frontier for Diagnosis and Prognosis of Kidney Diseases, Int J Mol Sci, 26(17), 5679
40. El Fekih, R. et al. (2025) An Exosomal mRNA Urine Test for Detection and Risk Stratification of Human Kidney Transplant Rejection, Kidney Int Rep, 10(4): 1131-1142
41. Alhadid, Y. et al. (2025) Development of a novel qPCR test for monitoring immunosuppression levels by quantifying torque teno virus (TTV) in whole blood, Human Immunology, 86 (5):111348
42. Kuczaj, A. et al. (2023) Torque Teno Virus (TTV) – A Potential Marker of Immunocompetence in Solid Organ Recipients, Viruses, 16 (1) 17
43. Haupenthal, F. et al. (2023) A multicentre, patient- and assessor-blinded, non-inferiority, randomised and controlled phase II trial to compare standard and torque teno virus-guided immunosuppression in kidney transplant recipients in the first year after transplantation: TTVguideIT, Trials, 24: 213
44. Feingold, B. et al. (2023) Short-term clinical outcomes and predicted cost savings of dd-cfDNA-led surveillance after pediatric heart transplantation, Clinical Transplantation, 37: e14933
45. Sharma, H. et al. (2024) The outcomes and Cost-Benefit Analysis of Cell-Free DNA (cfDNA) testing for organ transplant rejection, Abstract, BTS Annual Congress 2024 Abstract Book, P0027 p.166
Enquire about this article
"*" indicates required fields
