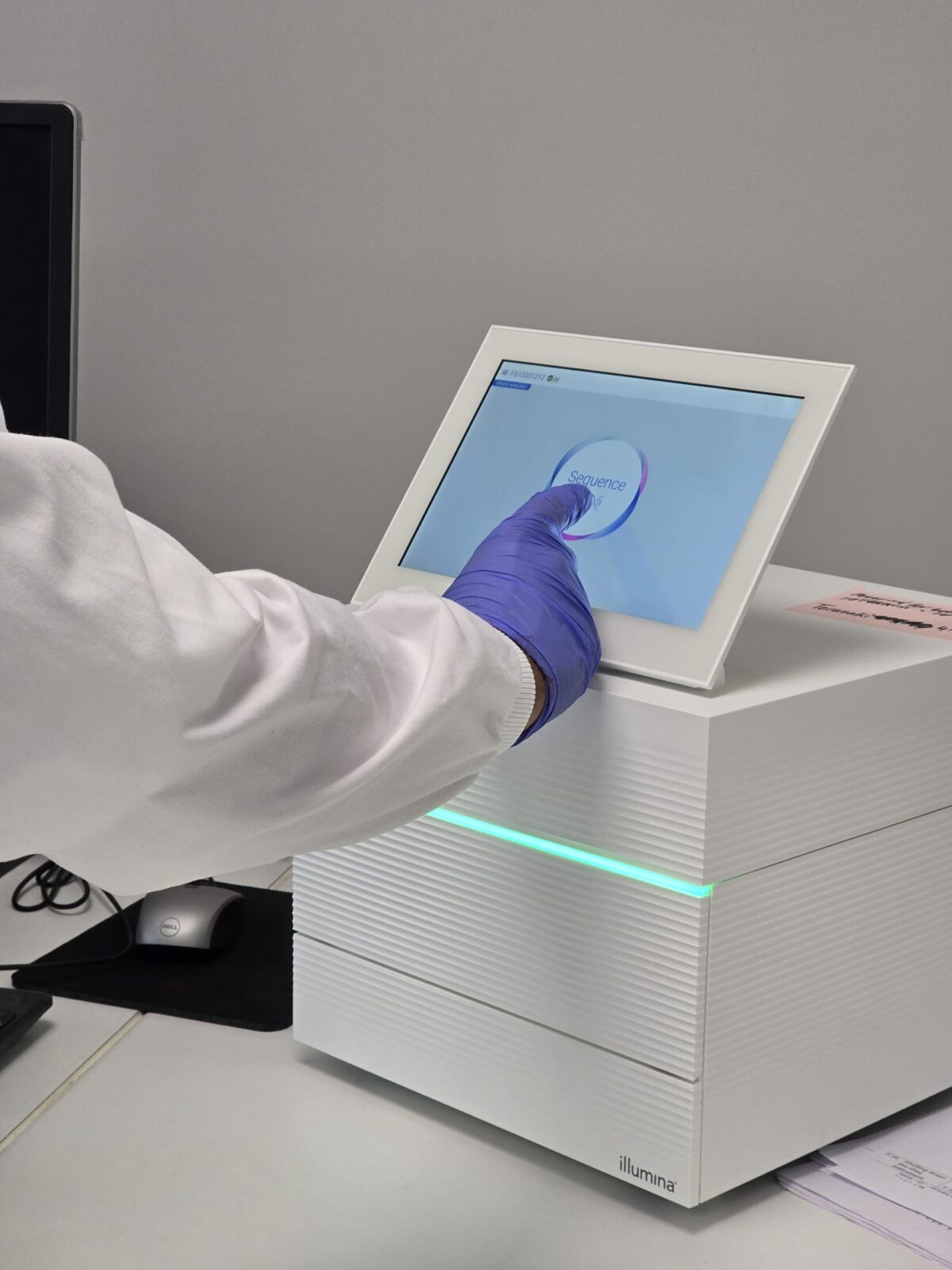
Read the analysis of the VH Bio team’s background using high-resolution HLA typing from buccal cell samples in this piece by Laboratory Scientist, Jakub:
When I started working at VH Bio in July 2023, I had only basic knowledge of Human Leukocyte Antigen (HLA) from my BSc degree. I understood the role of HLA in immunity and its importance in transplants. I was aware of HLA being extremely polymorphic, but I didn’t realise just how polymorphic HLA is until I started to see results obtained from our laboratory’s HLA Typing service. This sparked my interest, as I learned that a lot can be informed about a person’s heritage and background from their HLA result. Naturally, I was curious to see what my own HLA type could indicate about my background. I proposed the idea of HLA typing all our employees and there was a lot of excitement and enthusiasm about this, even from those without a scientific background. When starting the project, I asked everyone about their heritage and background to see if I can make links between this and their HLA results. This allowed me to get to know people on a more intimate level and I learned some historical lessons, too.
At VH Bio, we offer an intermediate-resolution HLA Typing service using a PCR-RSSO method (using One Lambda LABType SSO kits) and a high-resolution service using Next Generation Sequencing (NGS), (using One Lambda AllType FASTplex kits). I started off my training with intermediate resolution but later expanded my training to high-resolution HLA typing. I soon appreciated the importance of high-resolution genotyping because it gives largely unambiguous HLA results to three fields, whereas intermediate resolution HLA typing is ambiguous in the second field. I decided to use high-resolution HLA genotyping for this project in order to receive minimally ambiguous results. I utilised buccal swabs provided by Isohelix for sample collection and extracted DNA from these using the Maxwell RSC Genomic DNA Kit from Promega. NGS requires DNA of a very high quality and the DNA from the buccal swabs worked exceptionally well in our assay, portraying how robust the One Lambda AllType FASTplex kit is.
Analysing the results from the project confirmed the earlier statement with regards to HLA polymorphism; VH Bio’s cohort of employees was very polymorphic. A*03:01:01 was the most frequent of the HLA-A alleles, with seven individuals expressing this allele (Figure 1a). Before the project, I would have guessed that A*02:01:01 would be the most common HLA- A allele to be found in our company cohort because it’s the most common HLA-A allele worldwide with a frequency of approximately 50%.
The HLA-DPA1 locus is the least polymorphic HLA locus; this was confirmed in the project as 75% of all HLA-DPA1* alleles were DPA1*01:03:01 (Figure 1b).
Aside from these two loci, the other classical HLA loci (HLA-B, C, DRB1, DRB3/4/5, DQA1, DQB1, DPB1) were extremely polymorphic in our employee cohort, with no alleles standing out above any other as more common.
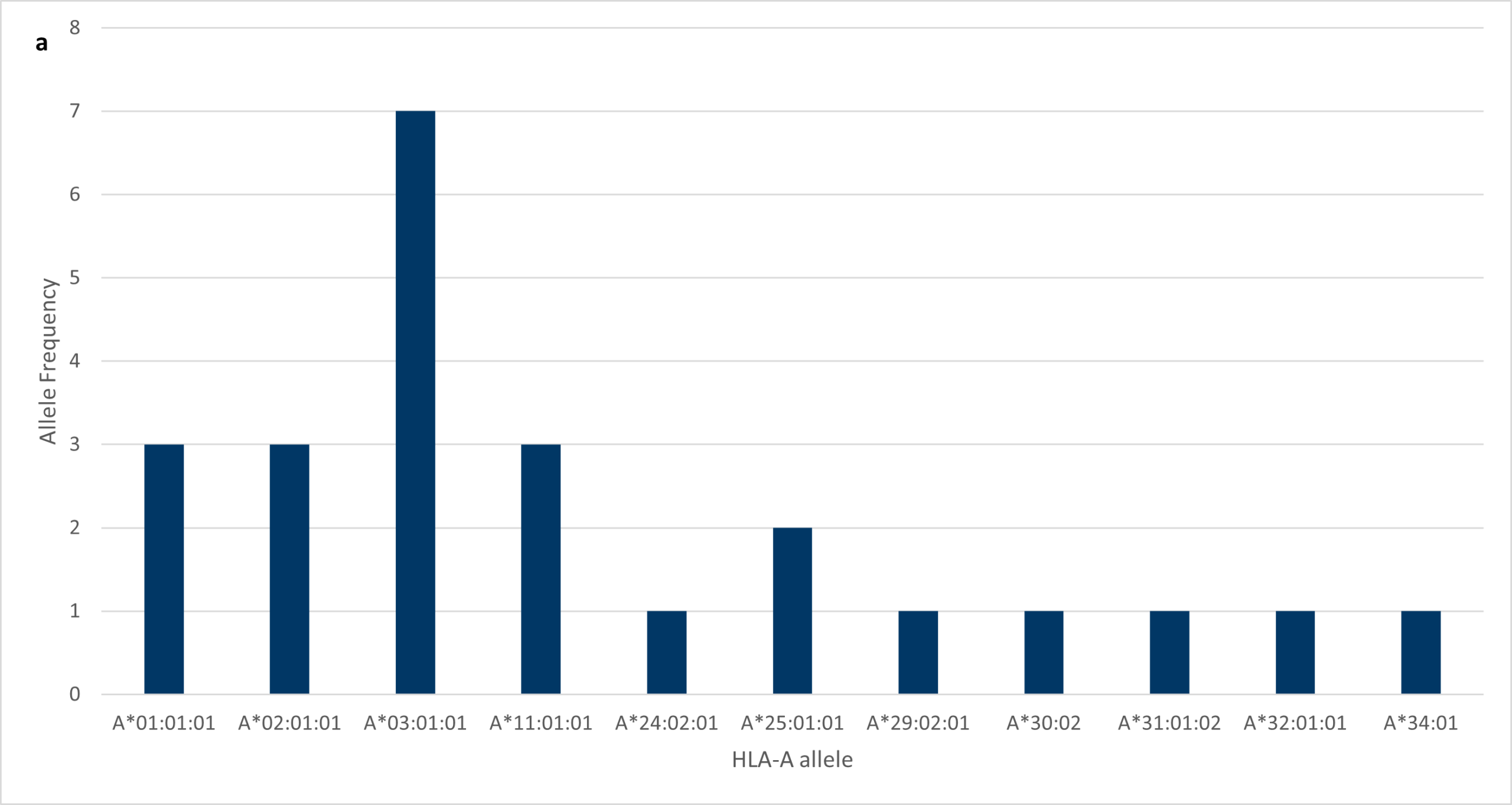
Figure 1: Data obtained from high-resolution HLA Genotyping using NGS. Figure 1a Frequency of HLA-A alleles amongst VH Bio employees.
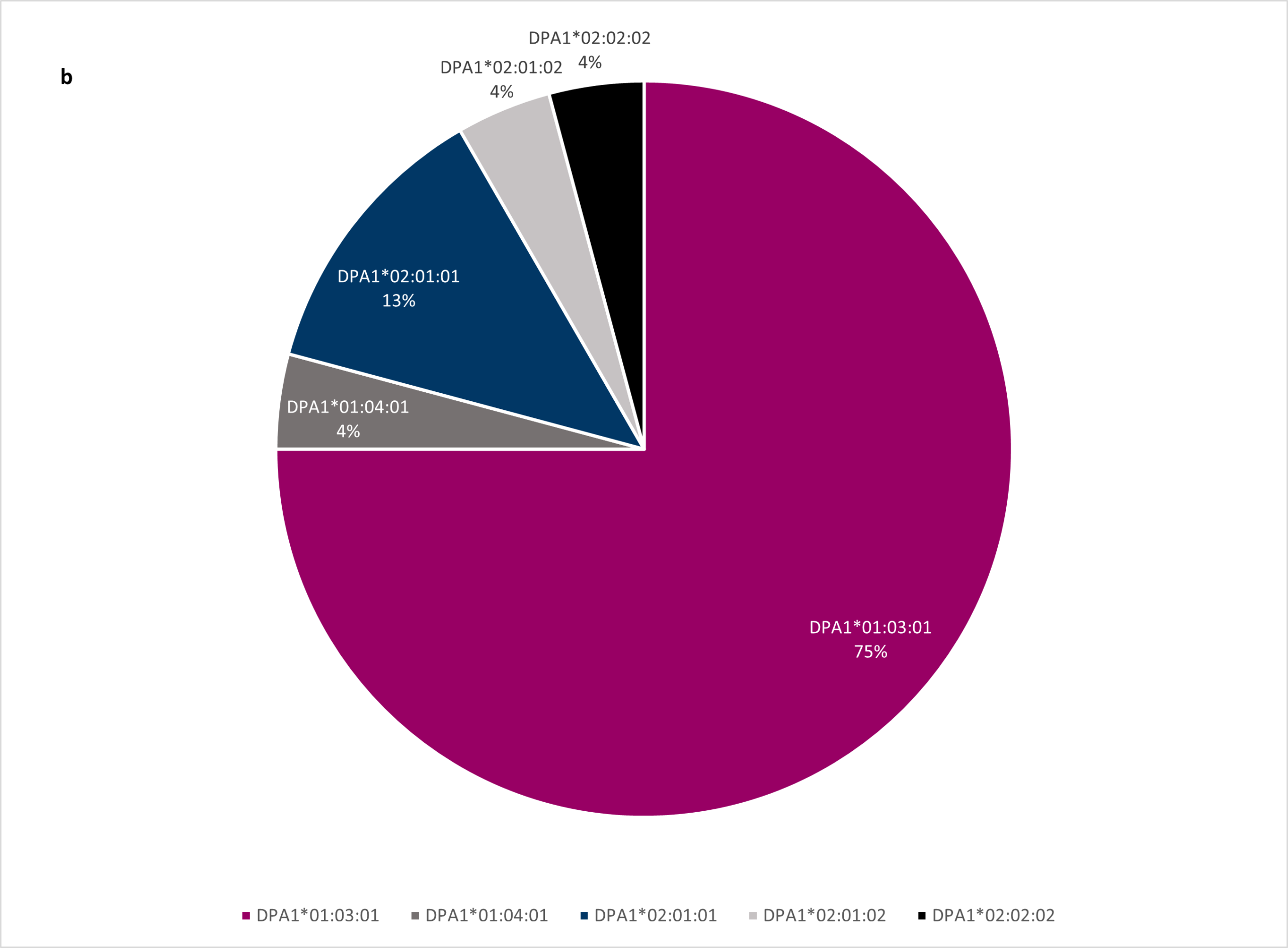
Figure 1b Percentage frequency of HLA-DPA1 alleles amongst VH Bio employees.
The HLA typing results obtained in the project provided interesting insight into the background of my colleagues. Jess Brookes’ mother originates from Fiji with her great grandmother being indigenous to Fiji and her great grandfather being Chinese, whilst father’s side originates from Northern Ireland. From Jess’ HLA Typing results we were able to see evidence of the Asian/Pacific background. HLA-A*34:01, B*13:01 and DRB1*15:02 are all commonly found in Asian/Pacific populations, A*34:01 and B*13:01 are rare in Caucasian populations. Using the data obtained we were able to construct the haplotypes which she likely obtained from her mother’s side and father’s side.
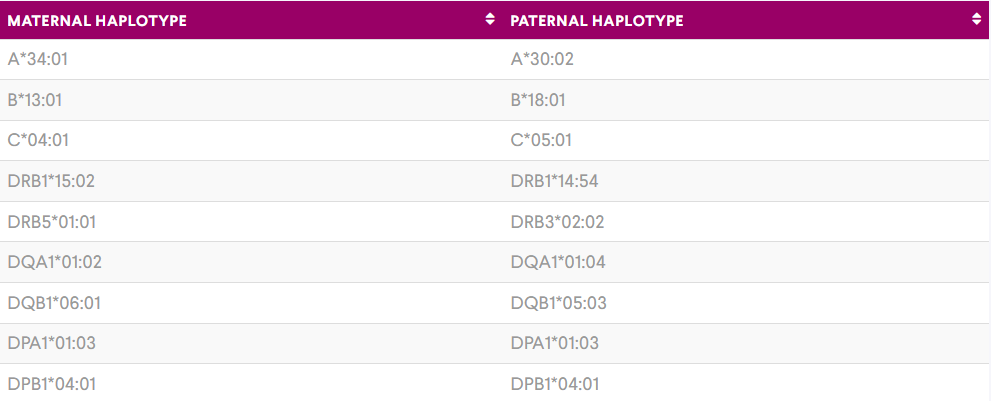
I completed a haplotype search on allelefrequencies.net, for both these assumed haplotypes and no results were returned, illustrating again the extent of HLA polymorphism. However, this could be partly due to the limitations of allelefrequencies.net that there is a lack of data from small countries such as Fiji who don’t have an established transplant program and therefore not large scale HLA typing within the population. Recently at VH Bio, we helped to set up a HLA typing programme in Barbados.
Jess’ results are a good example of why I decided to complete this project using high-resolution HLA genotyping rather than intermediate resolution. In intermediate-resolution typing, the analysis software does not distinguish between HLA-B*13:01 and B*13:02, whereas high-resolution typing provides an unambiguous result between these alleles. Similarly, HLA-DRB1*14:54 and DRB1*14:01 are not distinguished by intermediate-resolution typing. HLA-DRB1*14:54 is the most common HLA-DRB1*14 allele, which is a nice example that the most common HLA alleles are not always those with the lowest numerical value at the second field.
There was another example of evidence of heritage from HLA typing results when looking at Ali Rafique’s results. Ali has a Pakistani/Indian background; he has HLA-C*07:06:01 and HLA-DRB1*12:02:01 alleles, which are both extremely common in Asian populations. The advantage of high-resolution HLA typing was again shown here because intermediate-resolution typing doesn’t distinguish C*07:06 from the significantly more common C*07:01 allele. When speaking to Ali about his heritage I ended up with a little history lesson as his parents are Pakistani, but his grandparents are technically Indian because Pakistan didn’t gain independence from India until 1947.
Ali Rafique’s and Jess’ results were expected because we knew about their Asian and Asian/Pacific backgrounds. However, we came across some results that we did not expect. Pete has a very local background, as his parents and grandparents were all brought up around the Durham area. All of Pete’s HLA alleles were relatively commonly found in Caucasian populations apart from a surprising HLA-B*27:01, which is an extremely rare allele. After completing a search on allelefrequencies.net we discovered that this allele has been previously found in Iranian and Indian populations. Pete was very shocked to find out he had such a rare allele, and he came out with very interesting clinical questions asking about what implications this rare allele has if he ever needed a solid organ or stem cell transplant, and he was curious if he’s likely passed this rare allele down to his daughter.
Ben’s HLA Typing results contained a relatively uncommon DRB1*13:03 allele. His grandparents from his mother’s side come from Belgium and Germany, meanwhile from his father’s side they come from Northeast England and Sheffield. During Ben’s previous job in a clinical HLA lab, he ran his results through a bone marrow donor registry and had close matches in Eastern Europe.
Obviously, I was most interested in my own HLA type! I was extremely curious to see if I’ve got any rare or unexpected alleles which would teach me something surprising about my heritage. I was mildly disappointed to find out that my HLA alleles are relatively common in Caucasian populations. I come from a Polish background with both sets of parents and grandparents being Polish. However, something interesting that I learned from my HLA results is that my B*51:01:01 allele is prevalent across the historical Silk Road which connected the western world with the Middle East and Europe.

HLA is also an aid to diagnosis of and potentially associated with many diseases, and we saw some examples of these alleles in our VH Bio employee cohort. The best example is HLA-B*27 which is heavily associated with the autoimmune disease Ankylosing Spondylitis (AS), 90% of AS cases come from people with the HLA-B27 gene. This is one of many HLA genes which are associated with disease, other examples are Coeliac Disease, Type 1 Diabetes Mellitus, Rheumatoid Arthritis and many others. Personally, when looking at my HLA results the first locus I looked at was HLA-DQB1* because of the association of DQ2 and DQ8 with Coeliac Disease. I was relieved to see that I can continue to eat normal pizza and pasta containing gluten for the rest of my life!
The project has been both interesting and insightful on many different levels. From a scientific perspective, it has proved we can obtain high-resolution HLA typing from buccal cell samples, highlighted the advantages of high-resolution typing and the incredible level of HLA polymorphism which never stops making it interesting. From a team perspective it has allowed me to get to know my colleagues on a different level, learning about their backgrounds/cultures, leading to a few history lessons. From an education perspective it has brought scientific and non-scientific VH Bio employees closer.
VH Bio offers a Research Use Only intermediate-resolution HLA Typing service, providing results to two-fields, with ambiguities in the second field, and a high-resolution HLA Typing service with minimal ambiguous results to three-fields.
Enquire about this article
"*" indicates required fields
